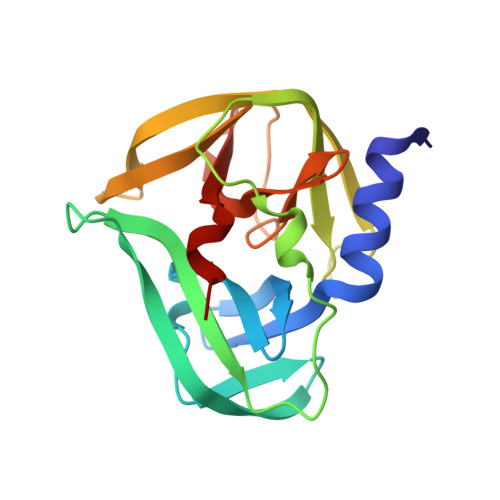
Crystallographic Fragment Screen of Coxsackievirus A16 2A Protease identifies new opportunities for the development of broad-spectrum anti-enterovirals.
Lithgo, R.M., Tomlinson, C.W.E., Fairhead, M., Winokan, M., Thompson, W., Wild, C., Aschenbrenner, J.C., Balcomb, B.H., Marples, P.G., Chandran, A.V., Golding, M., Koekemoer, L., Williams, E.P., Wang, S., Ni, X., MacLean, E., Giroud, C., Godoy, A.S., Xavier, M.A., Walsh, M., Fearon, D., von Delft, F.(2024) bioRxiv
- PubMed: 38746446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2024.04.29.591684
- Primary Citation Related Structures:
7GNV, 7GNW, 7GNX, 7GNY, 7GNZ, 7GO0, 7GO1, 7GO2, 7GO3, 7GO4, 7GO5, 7GO6, 7GO7, 7GO8, 7GO9, 7GOA, 7GOB, 7GOC, 7GOD, 7GOE, 7GOF, 7GOG, 7GOH, 7GOI, 7GOJ, 7GOK, 7GOL, 7GOM, 7GON, 7GOO, 7GOP, 7GOQ, 7GOR, 7GOS, 7GOT, 7GOU, 7GOV, 7GOW, 7GOX, 7GOY, 7GOZ, 7GP0, 7GP1, 7GP2, 7GP3, 7GP4, 7GP5, 7GP6, 7GP7, 7GP8, ... Search all related entries - PubMed Abstract:
Enteroviruses are the causative agents of paediatric hand-foot-and-mouth disease, and a target for pandemic preparedness due to the risk of higher order complications in a large-scale outbreak. The 2A protease of these viruses is responsible for the self-cleavage of the poly protein, allowing for correct folding and assembly of capsid proteins in the final stages of viral replication. These 2A proteases are highly conserved between Enterovirus species, such as Enterovirus A71 and Coxsackievirus A16 . Inhibition of the 2A protease deranges capsid folding and assembly, preventing formation of mature virions in host cells and making the protease a valuable target for antiviral activity. Herein, we describe a crystallographic fragment screening campaign that identified 75 fragments which bind to the 2A protease including 38 unique compounds shown to bind within the active site. These fragments reveal a path for the development of non-peptidomimetic inhibitors of the 2A protease with broad-spectrum anti-enteroviral activity.