ATP induced conformational changes facilitate E1-E2 disulfide bridging in the ubiquitin system.
Misra, M., Schaefer, A., Kuhn, M., Pluska, L., Tessmer, I., Sommer, T., Schindelin, H.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin-activating enzyme E1 1 | 1,024 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: UBA1, YKL210W EC: 6.2.1.45 | 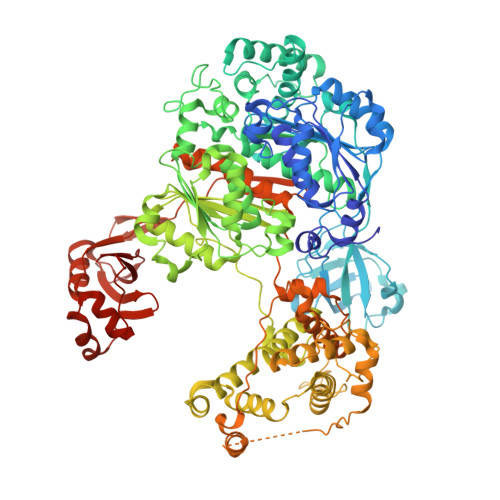 | |
UniProt | |||||
Find proteins for P22515 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P22515 Go to UniProtKB: P22515 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P22515 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin-conjugating enzyme E2 13 | 154 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: UBC13, YDR092W, YD6652.04 EC: 2.3.2.23 | 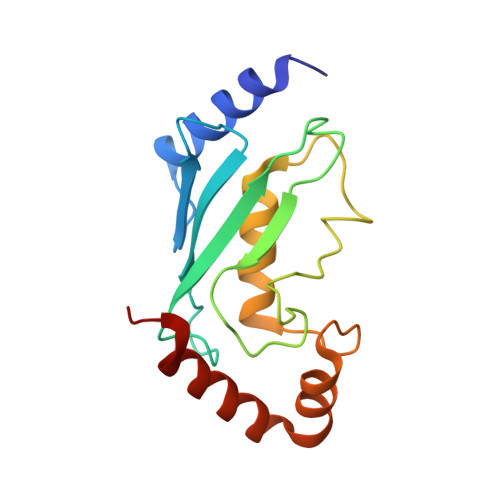 | |
UniProt | |||||
Find proteins for P52490 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P52490 Go to UniProtKB: P52490 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P52490 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Query on SO4 | D [auth A], E [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Query on GOL | F [auth A] G [auth A] H [auth A] I [auth A] J [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 177.71 | α = 90 |
| b = 72.9 | β = 112.43 |
| c = 139.52 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| Aimless | data scaling |
| XDS | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| German Research Foundation (DFG) | Germany | SCHI 425-6/1 |