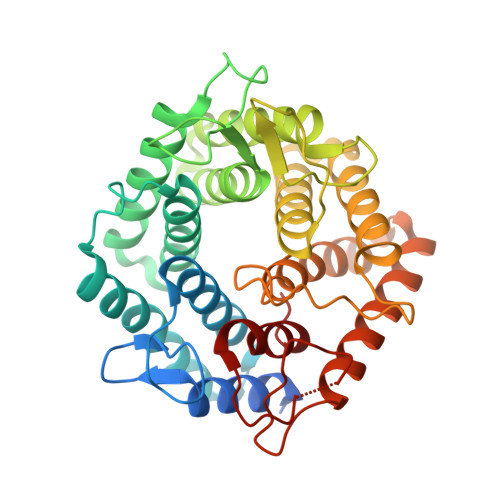
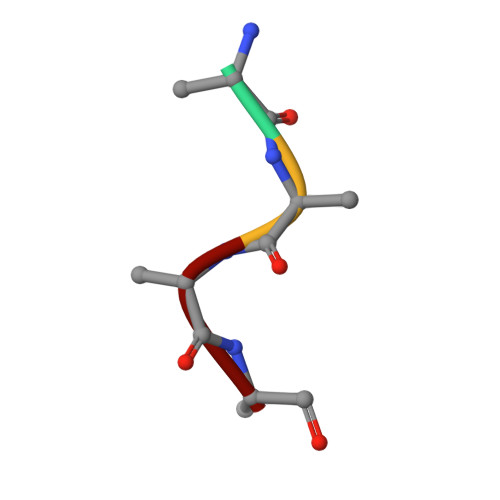
Development of Non-Hydrolysable Oligosaccharide Activity-Based Inactivators for Endoglycanases: A Case Study on alpha-1,6 Mannanases.
Schroder, S.P., Offen, W.A., Males, A., Jin, Y., de Boer, C., Enotarpi, J., Marino, L., van der Marel, G.A., Florea, B.I., Codee, J.D.C., Overkleeft, H.S., Davies, G.J.(2021) Chemistry 27: 9519-9523
- PubMed: 33878235 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/chem.202101255
- Primary Citation Related Structures:
6ZBM, 6ZBW, 6ZBX, 7NL5 - PubMed Abstract:
There is a vast genomic resource for enzymes active on carbohydrates. Lagging far behind, however, are functional chemical tools for the rapid characterization of carbohydrate-active enzymes. Activity-based probes (ABPs) offer one chemical solution to these issues with ABPs based upon cyclophellitol epoxide and aziridine covalent and irreversible inhibitors representing a potent and widespread approach. Such inhibitors for enzymes active on polysaccharides are potentially limited by the requirement for several glycosidic bonds, themselves substrates for the enzyme targets. Here, it is shown that non-hydrolysable trisaccharide can be synthesized and applied even to enzymes with challenging subsite requirements. It was found that incorporation of carbasugar moieties, which was accomplished by cuprate-assisted regioselective trans-diaxial epoxide opening of carba-mannal synthesised for this purpose, yields inactivators that act as powerful activity-based inhibitors for α-1,6 endo-mannanases. 3-D structures at 1.35-1.47 Å resolutions confirm the design rationale and binding to the enzymatic nucleophile. Carbasugar oligosaccharide cyclophellitols offer a powerful new approach for the design of robust endoglycosidase inhibitors, while the synthesis procedures presented here should allow adaptation towards activity-based endoglycosidase probes as well as configurational isosteres targeting other endoglycosidase families.
- Leiden Institute of Chemistry, Leiden University Einsteinweg 55, 2333, CC Leiden, The Netherlands.
Organizational Affiliation: