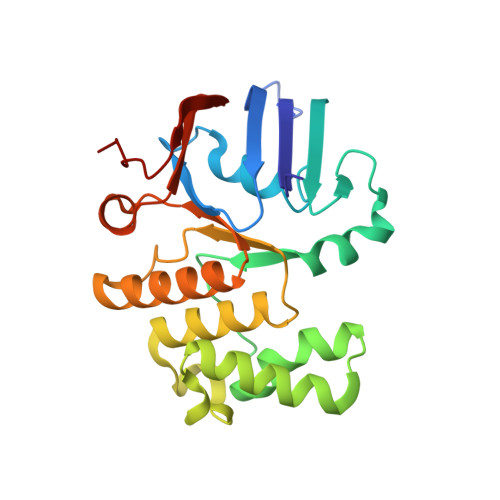
Structural Characterization of the Essential Cell Division Protein FtsE and Its Interaction with FtsX in Streptococcus pneumoniae.
Alcorlo, M., Straume, D., Lutkenhaus, J., Havarstein, L.S., Hermoso, J.A.(2020) mBio 11
- PubMed: 32873757 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.01488-20
- Primary Citation Related Structures:
6Z4W, 6Z63, 6Z67 - PubMed Abstract:
FtsEX is a membrane complex widely conserved across diverse bacterial genera and involved in critical processes such as recruitment of division proteins and in spatial and temporal regulation of muralytic activity during cell division or sporulation. FtsEX is a member of the ABC transporter superfamily. The component FtsX is an integral membrane protein, whereas FtsE is an ATPase and is required for the transmission of a conformational signal from the cytosol through the membrane to regulate the activity of cell wall hydrolases in the periplasm. Both proteins are essential in the major human respiratory pathogenic bacterium Streptococcus pneumoniae , and FtsX interacts with the modular peptidoglycan hydrolase PcsB at the septum. Here, we report high-resolution structures of pneumococcal FtsE bound to different nucleotides. Structural analysis revealed that FtsE contains all the conserved structural motifs associated with ATPase activity and afforded interpretation of the in vivo dimeric arrangement in both the ADP and ATP states. Interestingly, three specific FtsE regions with high structural plasticity were identified that shape the cavity in which the cytosolic region of FtsX would be inserted. The residues corresponding to the FtsX coupling helix, responsible for contacting FtsE, were identified and validated by in vivo mutagenesis studies showing that this interaction is essential for cell growth and proper morphology. IMPORTANCE Bacterial cell division is a central process that requires exquisite orchestration of both the cell wall biosynthetic and lytic machineries. The essential membrane complex FtsEX, widely conserved across bacteria, plays a central role by recruiting proteins to the divisome apparatus and by regulating periplasmic muralytic activity from the cytosol. FtsEX is a member of the type VII family of the ABC-superfamily, but instead of being a transporter, it couples the ATP hydrolysis catalyzed by FtsE to mechanically transduce a conformational signal that provokes the activation of peptidoglycan (PG) hydrolases. So far, no structural information is available for FtsE. Here, we provide the structural characterization of FtsE, confirming its ATPase nature and revealing regions with high structural plasticity which are key for FtsE binding to FtsX. The complementary binding region in FtsX has also been identified and validated in vivo Our results provide evidence on how the difference between the ATP/ADP-bound states in FtsE would dramatically alter the interaction of FtsEX with the PG hydrolase PcsB in pneumococcal division.
- Department of Crystallography and Structural Biology, Institute of Physical-Chemistry "Rocasolano", Spanish National Research Council (CSIC), Madrid, Spain.
Organizational Affiliation: