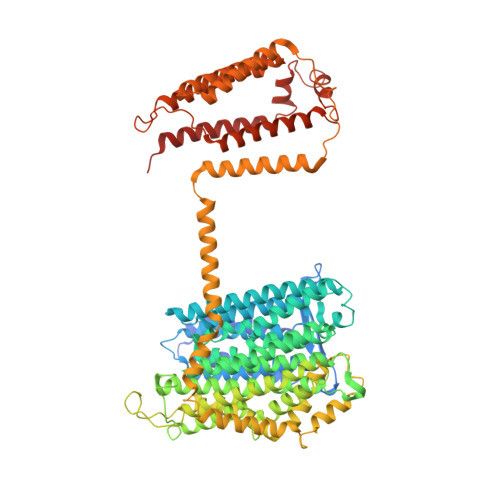
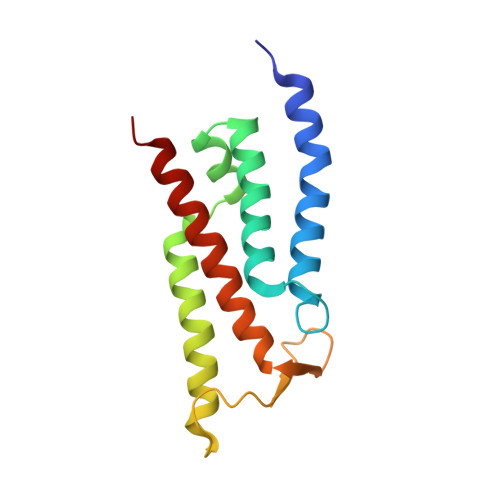
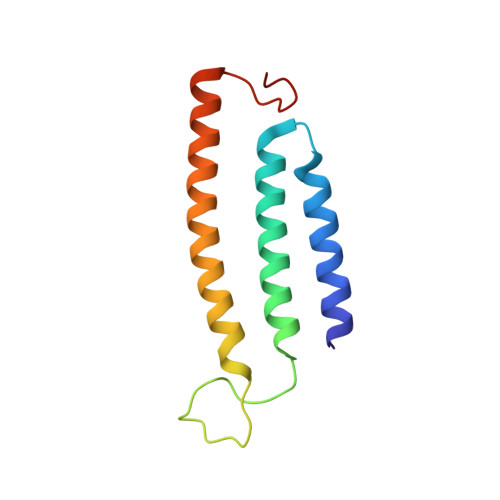
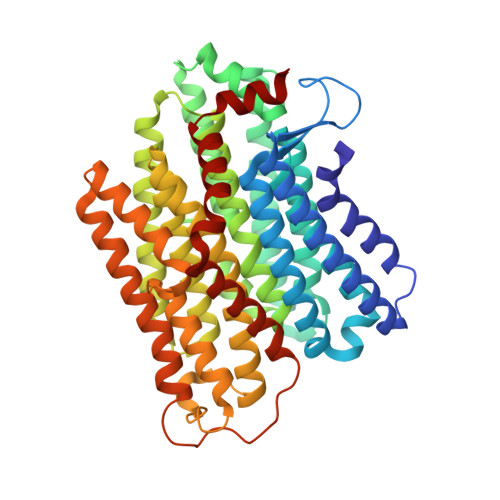
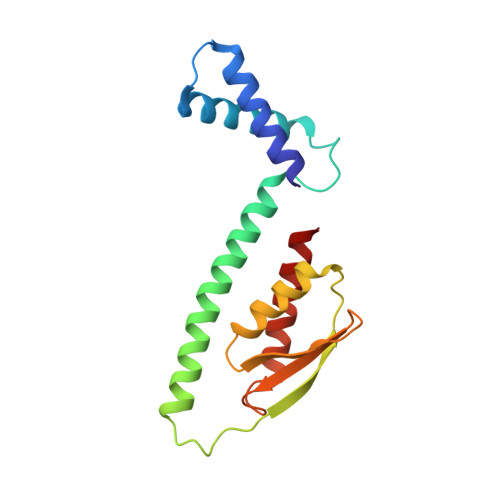
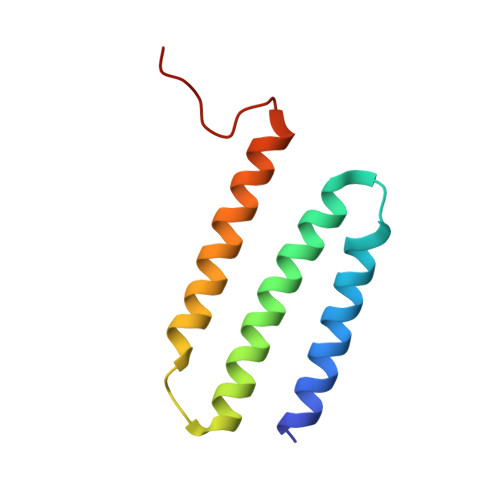
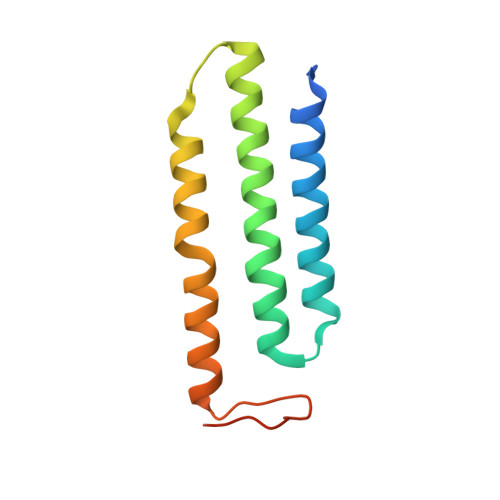
Structure and mechanism of the Mrp complex, an ancient cation/proton antiporter.
Steiner, J., Sazanov, L.(2020) Elife 9
- PubMed: 32735215 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.59407
- Primary Citation Related Structures:
6Z16 - PubMed Abstract:
Multiple resistance and pH adaptation (Mrp) antiporters are multi-subunit Na + (or K + )/H + exchangers representing an ancestor of many essential redox-driven proton pumps, such as respiratory complex I. The mechanism of coupling between ion or electron transfer and proton translocation in this large protein family is unknown. Here, we present the structure of the Mrp complex from Anoxybacillus flavithermus solved by cryo-EM at 3.0 Å resolution. It is a dimer of seven-subunit protomers with 50 trans-membrane helices each. Surface charge distribution within each monomer is remarkably asymmetric, revealing probable proton and sodium translocation pathways. On the basis of the structure we propose a mechanism where the coupling between sodium and proton translocation is facilitated by a series of electrostatic interactions between a cation and key charged residues. This mechanism is likely to be applicable to the entire family of redox proton pumps, where electron transfer to substrates replaces cation movements.
- Institute of Science and Technology Austria, Klosterneuburg, Austria.
Organizational Affiliation: