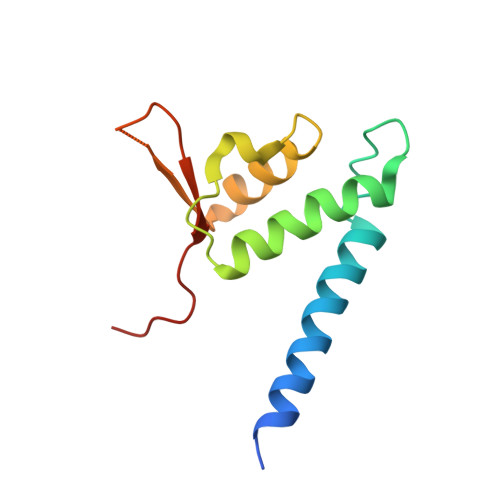
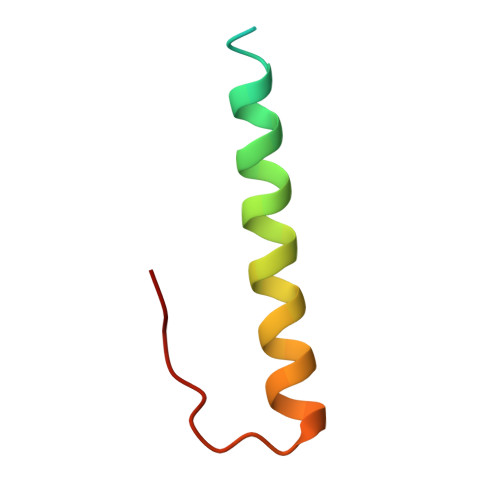
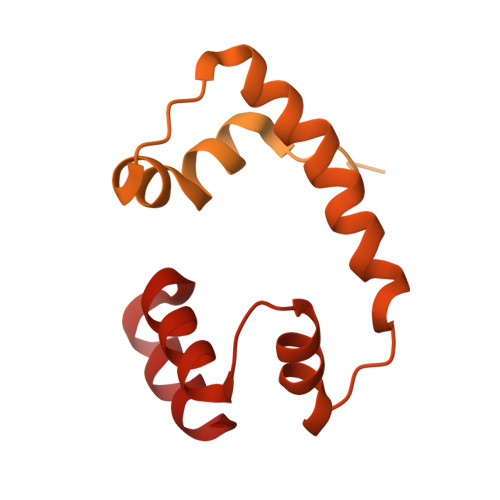
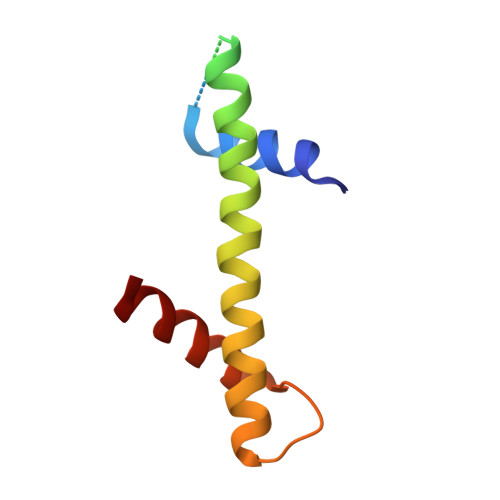
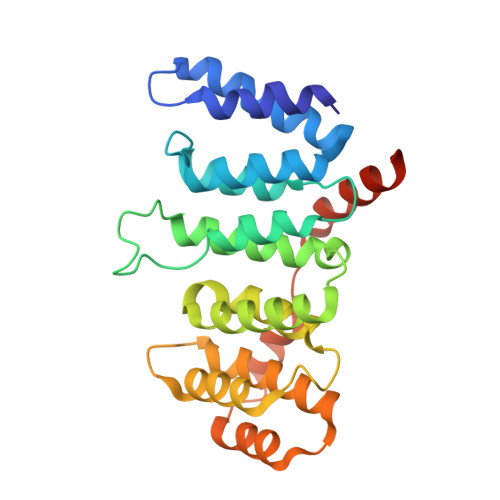
Crystal structure of the Cenp-HIKHead-TW sub-module of the inner kinetochore CCAN complex.
Zhang, Z., Bellini, D., Barford, D.(2020) Nucleic Acids Res 48: 11172-11184
- PubMed: 32976599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa772
- Primary Citation Related Structures:
6YPC - PubMed Abstract:
Kinetochores are large multi-subunit complexes that attach centromeric chromatin to microtubules of the mitotic spindle, enabling sister chromatid segregation in mitosis. The inner kinetochore constitutive centromere associated network (CCAN) complex assembles onto the centromere-specific Cenp-A nucleosome (Cenp-ANuc), thereby coupling the centromere to the microtubule-binding outer kinetochore. CCAN is a conserved 14-16 subunit complex composed of discrete modules. Here, we determined the crystal structure of the Saccharomyces cerevisiae Cenp-HIKHead-TW sub-module, revealing how Cenp-HIK and Cenp-TW interact at the conserved Cenp-HIKHead-Cenp-TW interface. A major interface is formed by the C-terminal anti-parallel α-helices of the histone fold extension (HFE) of the Cenp-T histone fold domain (HFD) combining with α-helix H3 of Cenp-K to create a compact three α-helical bundle. We fitted the Cenp-HIKHead-TW sub-module to the previously determined cryo-EM map of the S. cerevisiae CCAN-Cenp-ANuc complex. This showed that the HEAT repeat domain of Cenp-IHead and C-terminal HFD of Cenp-T of the Cenp-HIKHead-TW sub-module interact with the nucleosome DNA gyre at a site close to the Cenp-ANuc dyad axis. Our structure provides a framework for understanding how Cenp-T links centromeric Cenp-ANuc to the outer kinetochore through its HFD and N-terminal Ndc80-binding motif, respectively.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: