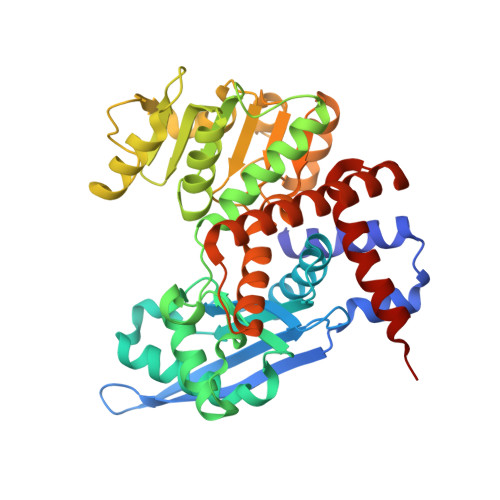
Structural Studies of Glutamate Dehydrogenase (Isoform 1) FromArabidopsis thaliana, an Important Enzyme at the Branch-Point Between Carbon and Nitrogen Metabolism.
Grzechowiak, M., Sliwiak, J., Jaskolski, M., Ruszkowski, M.(2020) Front Plant Sci 11: 754-754
- PubMed: 32655590
- DOI: https://doi.org/10.3389/fpls.2020.00754
- Primary Citation Related Structures:
6YEH, 6YEI - PubMed Abstract:
Glutamate dehydrogenase (GDH) releases ammonia in a reversible NAD(P) + -dependent oxidative deamination of glutamate that yields 2-oxoglutarate (2OG). In current perception, GDH contributes to Glu homeostasis and plays a significant role at the junction of carbon and nitrogen assimilation pathways. GDHs are members of a superfamily of ELFV (Glu/Leu/Phe/Val) amino acid dehydrogenases and are subdivided into three subclasses, based on coenzyme specificity: NAD + -specific, NAD + /NADP + dual-specific, and NADP + -specific. We determined in this work that the mitochondrial At GDH1 isozyme from A. thaliana is NAD + -specific. Altogether, A. thaliana expresses three GDH isozymes ( At GDH1-3) targeted to mitochondria, of which At GDH2 has an extra EF-hand motif and is stimulated by calcium. Our enzymatic assays of At GDH1 established that its sensitivity to calcium is negligible. In vivo the At GDH1-3 enzymes form homo- and heterohexamers of varied composition. We solved the crystal structure of recombinant At GDH1 in the apo-form and in complex with NAD + at 2.59 and 2.03 Å resolution, respectively. We demonstrate also that both in the apo form and in 1:1 complex with NAD + , it forms D 3 -symmetric homohexamers. A subunit of At GDH1 consists of domain I, which is involved in hexamer formation and substrate binding, and of domain II which binds coenzyme. Most of the subunits in our crystal structures, including those in NAD + complex, are in open conformation, with domain II forming a large (albeit variable) angle with domain I. One of the subunits of the At GDH1-NAD + hexamer contains a serendipitous 2OG molecule in the active site, causing a dramatic (∼25°) closure of the domains. We provide convincing evidence that the N-terminal peptide preceding domain I is a mitochondrial targeting signal, with a predicted cleavage site for mitochondrial processing peptidase (MPP) at Leu17-Leu18 that is followed by an unexpected potassium coordination site (Ser27, Ile30). We also identified several MPD [(+/-)-2-methyl-2,4-pentanediol] binding sites with conserved sequence. Although At GDH1 is insensitive to MPD in our assays, the observation of druggable sites opens a potential for non-competitive herbicide design.
- Center for Biocrystallographic Research Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznań, Poland.
Organizational Affiliation: