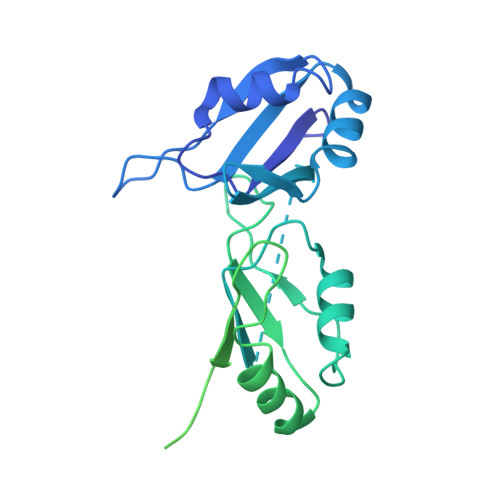
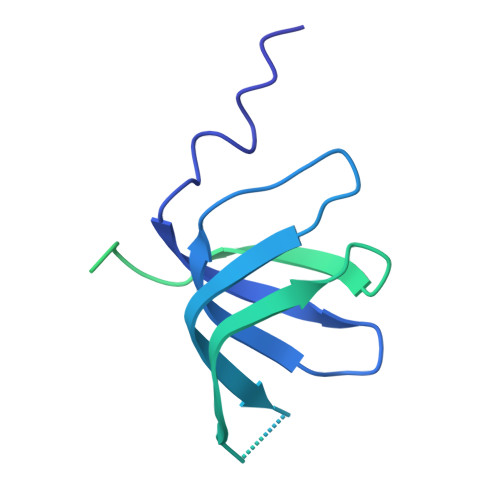
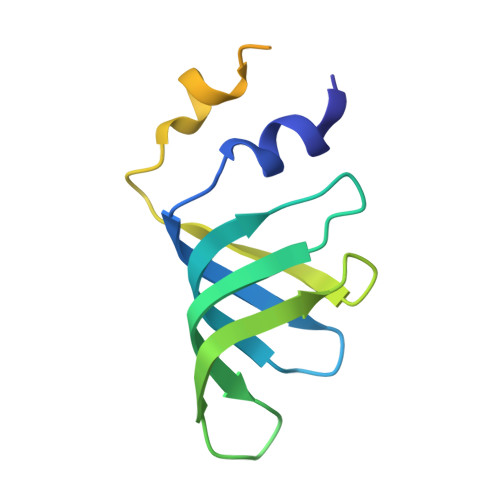
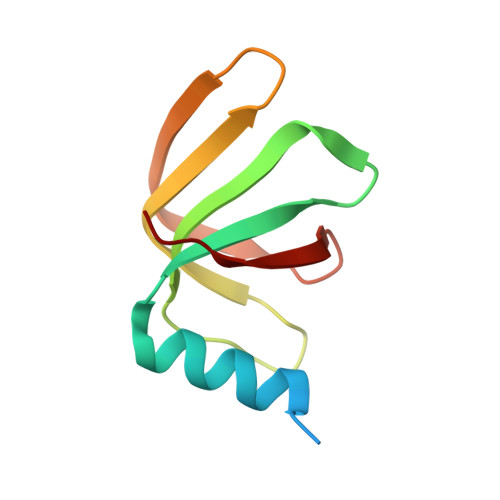
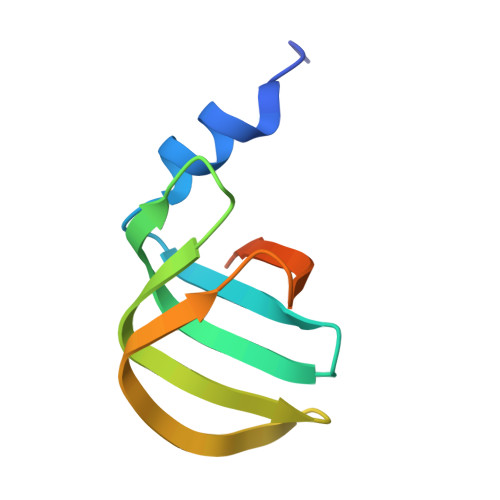
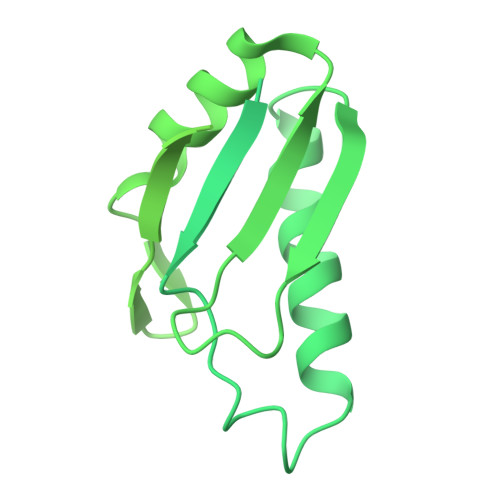
Molecular architecture of the human 17S U2 snRNP.
Zhang, Z., Will, C.L., Bertram, K., Dybkov, O., Hartmuth, K., Agafonov, D.E., Hofele, R., Urlaub, H., Kastner, B., Luhrmann, R., Stark, H.(2020) Nature 583: 310-313
- PubMed: 32494006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2344-3
- Primary Citation Related Structures:
6Y50, 6Y53, 6Y5Q - PubMed Abstract:
The U2 small nuclear ribonucleoprotein (snRNP) has an essential role in the selection of the precursor mRNA branch-site adenosine, the nucleophile for the first step of splicing 1 . Stable addition of U2 during early spliceosome formation requires the DEAD-box ATPase PRP5 2-7 . Yeast U2 small nuclear RNA (snRNA) nucleotides that form base pairs with the branch site are initially sequestered in a branchpoint-interacting stem-loop (BSL) 8 , but whether the human U2 snRNA folds in a similar manner is unknown. The U2 SF3B1 protein, a common mutational target in haematopoietic cancers 9 , contains a HEAT domain (SF3B1 HEAT ) with an open conformation in isolated SF3b 10 , but a closed conformation in spliceosomes 11 , which is required for stable interaction between U2 and the branch site. Here we report a 3D cryo-electron microscopy structure of the human 17S U2 snRNP at a core resolution of 4.1 Å and combine it with protein crosslinking data to determine the molecular architecture of this snRNP. Our structure reveals that SF3B1 HEAT interacts with PRP5 and TAT-SF1, and maintains its open conformation in U2 snRNP, and that U2 snRNA forms a BSL that is sandwiched between PRP5, TAT-SF1 and SF3B1 HEAT . Thus, substantial remodelling of the BSL and displacement of BSL-interacting proteins must occur to allow formation of the U2-branch-site helix. Our studies provide a structural explanation of why TAT-SF1 must be displaced before the stable addition of U2 to the spliceosome, and identify RNP rearrangements facilitated by PRP5 that are required for stable interaction between U2 and the branch site.
- Department of Structural Dynamics, MPI for Biophysical Chemistry, Göttingen, Germany.
Organizational Affiliation: