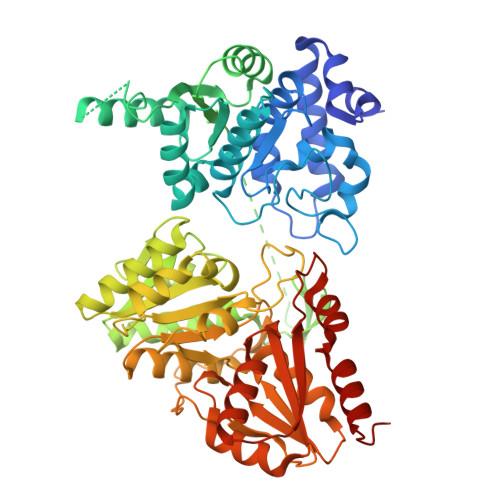
Identification of a 1-deoxy-D-xylulose-5-phosphate synthase (DXS) mutant with improved crystallographic properties.
Gierse, R.M., Reddem, E.R., Alhayek, A., Baitinger, D., Hamid, Z., Jakobi, H., Laber, B., Lange, G., Hirsch, A.K.H., Groves, M.R.(2021) Biochem Biophys Res Commun 539: 42-47
- PubMed: 33421767
- DOI: https://doi.org/10.1016/j.bbrc.2020.12.069
- Primary Citation of Related Structures:
6XXG - PubMed Abstract:
In this report, we describe a truncated Deinococcus radiodurans 1-deoxy-D-xylulose-5-phosphate synthase (DXS) protein that retains enzymatic activity, while slowing protein degradation and showing improved crystallization properties. With modern drug-design approaches relying heavily on the elucidation of atomic interactions of potential new drugs with their targets, the need for co-crystal structures with the compounds of interest is high. DXS itself is a promising drug target, as it catalyzes the first reaction in the 2-C-methyl-D-erythritol 4-phosphate (MEP)-pathway for the biosynthesis of the universal precursors of terpenes, which are essential secondary metabolites. In contrast to many bacteria and pathogens, which employ the MEP pathway, mammals use the distinct mevalonate-pathway for the biosynthesis of these precursors, which makes all enzymes of the MEP-pathway potential new targets for the development of anti-infectives. However, crystallization of DXS has proven to be challenging: while the first X-ray structures from Escherichia coli and D. radiodurans were solved in 2004, since then only two additions have been made in 2019 that were obtained under anoxic conditions. The presented site of truncation can potentially also be transferred to other homologues, opening up the possibility for the determination of crystal structures from pathogenic species, which until now could not be crystallized. This manuscript also provides a further example that truncation of a variable region of a protein can lead to improved structural data.
- Department for Drug Design and Optimization, Helmholtz Institute for Pharmaceutical Research (HIPS) - Helmholtz Centre for Infection Research (HZI), Campus Building E 8.1, 66123, Saarbrücken, Germany; Department of Pharmacy, Saarland University, Campus Building E8.1, 66123, Saarbrücken, Germany; Stratingh Institute for Chemistry, University of Groningen, Nijenborgh 7, 9747, AG Groningen, Netherlands.
Organizational Affiliation: