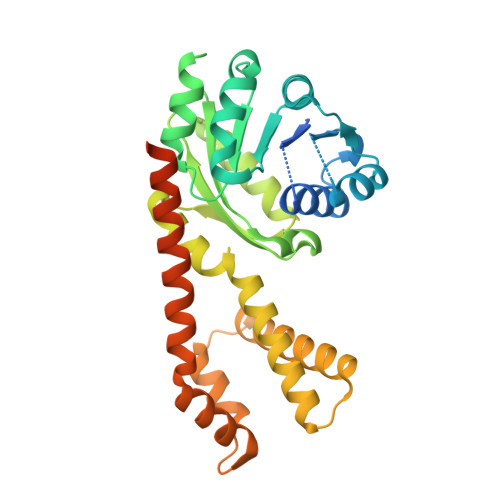
In crystallo screening for proline analog inhibitors of the proline cycle enzyme PYCR1.
Christensen, E.M., Bogner, A.N., Vandekeere, A., Tam, G.S., Patel, S.M., Becker, D.F., Fendt, S.M., Tanner, J.J.(2020) J Biological Chem 295: 18316-18327
- PubMed: 33109600
- DOI: https://doi.org/10.1074/jbc.RA120.016106
- Primary Citation Related Structures:
6XOZ, 6XP0, 6XP1, 6XP2, 6XP3 - PubMed Abstract:
Pyrroline-5-carboxylate reductase 1 (PYCR1) catalyzes the biosynthetic half-reaction of the proline cycle by reducing Δ 1 -pyrroline-5-carboxylate (P5C) to proline through the oxidation of NAD(P)H. Many cancers alter their proline metabolism by up-regulating the proline cycle and proline biosynthesis, and knockdowns of PYCR1 lead to decreased cell proliferation. Thus, evidence is growing for PYCR1 as a potential cancer therapy target. Inhibitors of cancer targets are useful as chemical probes for studying cancer mechanisms and starting compounds for drug discovery; however, there is a notable lack of validated inhibitors for PYCR1. To fill this gap, we performed a small-scale focused screen of proline analogs using X-ray crystallography. Five inhibitors of human PYCR1 were discovered: l-tetrahydro-2-furoic acid, cyclopentanecarboxylate, l-thiazolidine-4-carboxylate, l-thiazolidine-2-carboxylate, and N -formyl l-proline (NFLP). The most potent inhibitor was NFLP, which had a competitive (with P5C) inhibition constant of 100 μm The structure of PYCR1 complexed with NFLP shows that inhibitor binding is accompanied by conformational changes in the active site, including the translation of an α-helix by 1 Å. These changes are unique to NFLP and enable additional hydrogen bonds with the enzyme. NFLP was also shown to phenocopy the PYCR1 knockdown in MCF10A H-RAS V12 breast cancer cells by inhibiting de novo proline biosynthesis and impairing spheroidal growth. In summary, we generated the first validated chemical probe of PYCR1 and demonstrated proof-of-concept for screening proline analogs to discover inhibitors of the proline cycle.
- Department of Chemistry, University of Missouri, Columbia, Missouri, USA.
Organizational Affiliation: