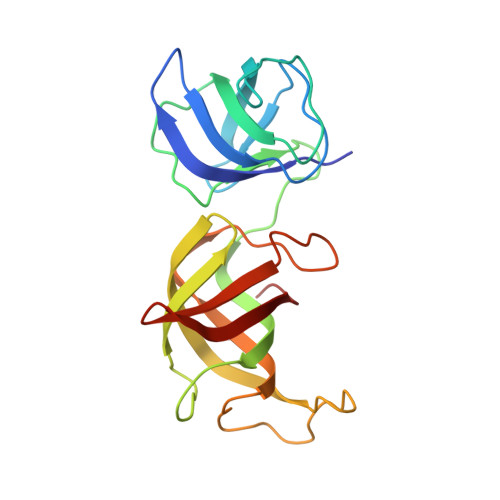
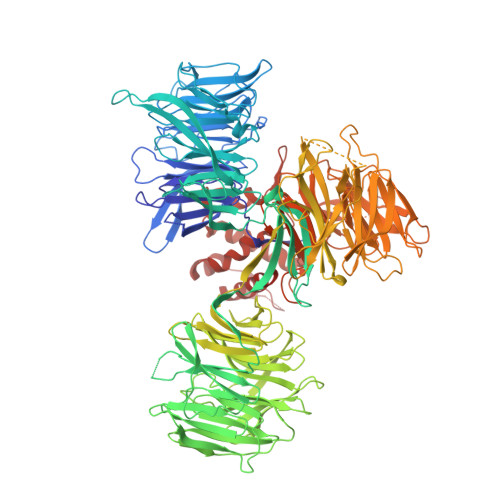
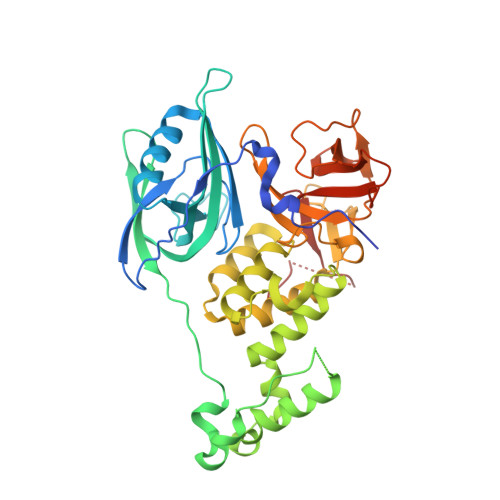
CC-90009, a novel cereblon E3 ligase modulator, targets acute myeloid leukemia blasts and leukemia stem cells.
Surka, C., Jin, L., Mbong, N., Lu, C.C., Jang, I.S., Rychak, E., Mendy, D., Clayton, T., Tindall, E., Hsu, C., Fontanillo, C., Tran, E., Contreras, A., Ng, S.W.K., Matyskiela, M., Wang, K., Chamberlain, P., Cathers, B., Carmichael, J., Hansen, J., Wang, J.C.Y., Minden, M.D., Fan, J., Pierce, D.W., Pourdehnad, M., Rolfe, M., Lopez-Girona, A., Dick, J.E., Lu, G.(2021) Blood 137: 661-677
- PubMed: 33197925
- DOI: https://doi.org/10.1182/blood.2020008676
- Primary Citation Related Structures:
6XK9 - PubMed Abstract:
A number of clinically validated drugs have been developed by repurposing the CUL4-DDB1-CRBN-RBX1 (CRL4CRBN) E3 ubiquitin ligase complex with molecular glue degraders to eliminate disease-driving proteins. Here, we present the identification of a first-in-class GSPT1-selective cereblon E3 ligase modulator, CC-90009. Biochemical, structural, and molecular characterization demonstrates that CC-90009 coopts the CRL4CRBN to selectively target GSPT1 for ubiquitination and proteasomal degradation. Depletion of GSPT1 by CC-90009 rapidly induces acute myeloid leukemia (AML) apoptosis, reducing leukemia engraftment and leukemia stem cells (LSCs) in large-scale primary patient xenografting of 35 independent AML samples, including those with adverse risk features. Using a genome-wide CRISPR-Cas9 screen for effectors of CC-90009 response, we uncovered the ILF2 and ILF3 heterodimeric complex as a novel regulator of cereblon expression. Knockout of ILF2/ILF3 decreases the production of full-length cereblon protein via modulating CRBN messenger RNA alternative splicing, leading to diminished response to CC-90009. The screen also revealed that the mTOR signaling and the integrated stress response specifically regulate the response to CC-90009 in contrast to other cereblon modulators. Hyperactivation of the mTOR pathway by inactivation of TSC1 and TSC2 protected against the growth inhibitory effect of CC-90009 by reducing CC-90009-induced binding of GSPT1 to cereblon and subsequent GSPT1 degradation. On the other hand, GSPT1 degradation promoted the activation of the GCN1/GCN2/ATF4 pathway and subsequent apoptosis in AML cells. Collectively, CC-90009 activity is mediated by multiple layers of signaling networks and pathways within AML blasts and LSCs, whose elucidation gives insight into further assessment of CC-90009s clinical utility. These trials were registered at www.clinicaltrials.gov as #NCT02848001 and #NCT04336982).
- Bristol-Myers Squibb, San Diego, CA.
Organizational Affiliation: