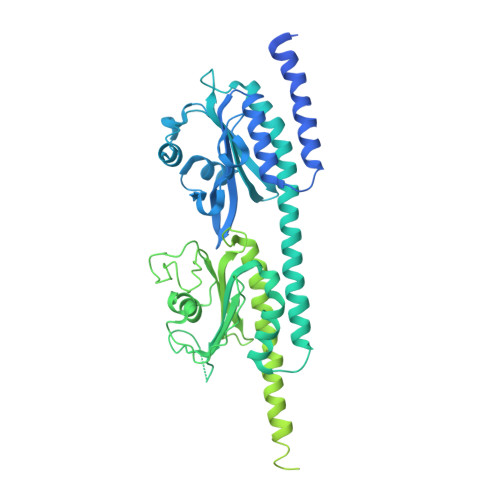
Structural Analysis of the Regulatory GAF Domains of cGMP Phosphodiesterase Elucidates the Allosteric Communication Pathway.
Gupta, R., Liu, Y., Wang, H., Nordyke, C.T., Puterbaugh, R.Z., Cui, W., Varga, K., Chu, F., Ke, H., Vashisth, H., Cote, R.H.(2020) J Mol Biology 432: 5765-5783
- PubMed: 32898583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2020.08.026
- Primary Citation Related Structures:
6X88, 9A0L, 9A0M, 9A0N, 9A0O - PubMed Abstract:
Regulation of photoreceptor phosphodiesterase (PDE6) activity is responsible for the speed, sensitivity, and recovery of the photoresponse during visual signaling in vertebrate photoreceptor cells. It is hypothesized that physiological differences in the light responsiveness of rods and cones may result in part from differences in the structure and regulation of the distinct isoforms of rod and cone PDE6. Although rod and cone PDE6 catalytic subunits share a similar domain organization consisting of tandem GAF domains (GAFa and GAFb) and a catalytic domain, cone PDE6 is a homodimer whereas rod PDE6 consists of two homologous catalytic subunits. Here we provide the x-ray crystal structure of cone GAFab regulatory domain solved at 3.3 Å resolution, in conjunction with chemical cross-linking and mass spectrometric analysis of conformational changes to GAFab induced upon binding of cGMP and the PDE6 inhibitory γ-subunit (Pγ). Ligand-induced changes in cross-linked residues implicate multiple conformational changes in the GAFa and GAFb domains in forming an allosteric communication network. Molecular dynamics simulations of cone GAFab revealed differences in conformational dynamics of the two subunits forming the homodimer and allosteric perturbations on cGMP binding. Cross-linking of Pγ to GAFab in conjunction with solution NMR spectroscopy of isotopically labeled Pγ identified the central polycationic region of Pγ interacting with the GAFb domain. These results provide a mechanistic basis for developing allosteric activators of PDE6 with therapeutic implications for halting the progression of several retinal degenerative diseases.
- Department of Molecular, Cellular and Biomedical Sciences, University of New Hampshire, 46 College Rd., Durham, NH 03824, USA.
Organizational Affiliation: