A Key Glycine in Bacterial Steroid-Degrading Acyl-CoA Dehydrogenases Allows Flavin-Ring Repositioning and Modulates Substrate Side Chain Specificity.
Stirling, A.J., Gilbert, S.E., Conner, M., Mallette, E., Kimber, M.S., Seah, S.Y.K.(2020) Biochemistry 59: 4081-4092
- PubMed: 33040522 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.0c00568
- Primary Citation Related Structures:
6WY8, 6WY9 - PubMed Abstract:
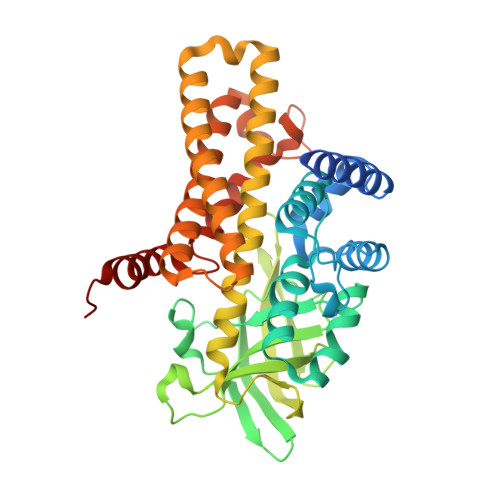
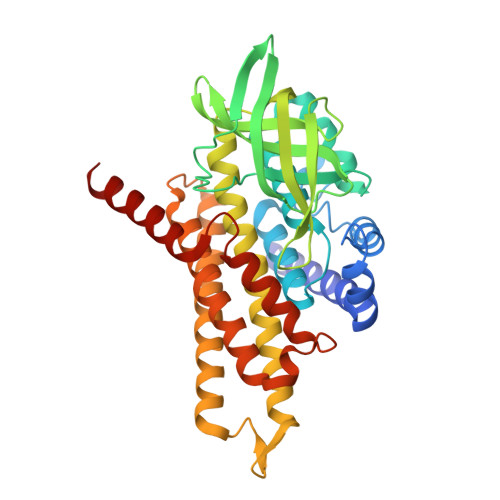
A wide variety of steroid metabolites synthesized by eukaryotes are all ultimately catabolized by bacteria; while generally saprophytic, pathogenic Mycobacteria have repurposed these pathways to utilize host intracellular cholesterol pools. Steroid degradation is complex, but a recurring theme is that cycles of β-oxidation are used to iteratively remove acetyl- or propanoyl-CoA groups. These β-oxidation cycles are initiated by the FAD-dependent oxidation of acyl groups, catalyzed by acyl-CoA dehydrogenases (ACADs). We show here that the tcur3481 and tcur3483 genes of Thermomonospora curvata encode subunits of a single ACAD that degrades steroid side chains with a preference for three-carbon over five-carbon substituents. The structure confirms that this enzyme is heterotetrameric, with active sites only in the Tcur3483 subunits. In comparison with the steroid ACAD FadE26-FadE27 from Mycobacterium tuberculosis , the active site is narrower and closed at the steroid-binding end, suggesting that Tcur3481-Tcur3483 is in a catalytically productive state, while FadE26-FadE27 is opened up to allow substrate entry. The flavin rings in Tcur3481-Tcur3483 sit in an unusual pocket created by Gly363, a residue conserved as Ala in steroid ACADs narrowly specific for five-carbon side chains, including FadE34. A Gly363Ala variant of Tcur3481-Tcur3483 prefers five-carbon side chains, while an inverse Ala691Gly FadE34 variant enables three-carbon side chain steroid oxidation. We determined the structure of the Tcur3483 Gly363Ala variant, showing that the flavin rings shift into the more conventional position. Modeling suggests that the shifted flavin position made possible by Gly363 is required to allow the bulky, inflexible three-carbon steroid to bind productively in the active site.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario, Canada N1G 5E9.
Organizational Affiliation: