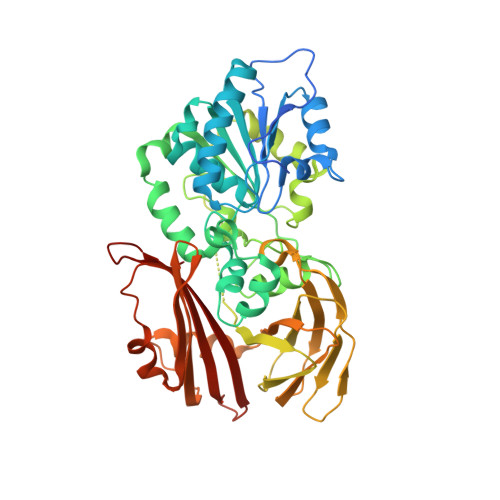
Structures of BlEst2 from Bacillus licheniformis in its propeptide and mature forms reveal autoinhibitory effects of the C-terminal domain
Nakamura, A.M., Godoy, A.S., Kadowaki, M.A.S., Trentin, L.N., Gonzalez, S.E.T., Skaf, M.S., Polikarpov, I.(2024) FEBS J n/a