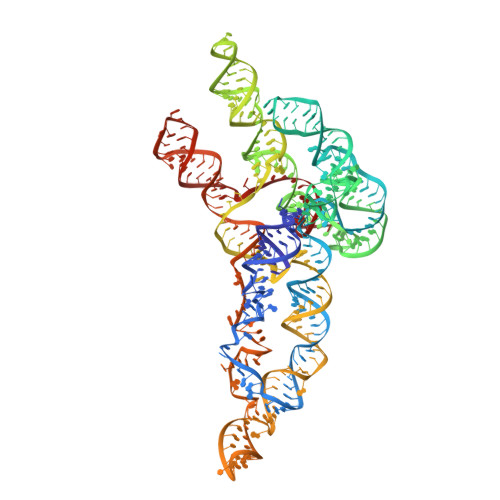
Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures.
Kappel, K., Zhang, K., Su, Z., Watkins, A.M., Kladwang, W., Li, S., Pintilie, G., Topkar, V.V., Rangan, R., Zheludev, I.N., Yesselman, J.D., Chiu, W., Das, R.(2020) Nat Methods 17: 699-707
- PubMed: 32616928
- DOI: https://doi.org/10.1038/s41592-020-0878-9
- Primary Citation of Related Structures:
6WLJ, 6WLK, 6WLL, 6WLM, 6WLN, 6WLO, 6WLQ, 6WLR, 6WLS, 6WLT, 6WLU - PubMed Abstract:
The discovery and design of biologically important RNA molecules is outpacing three-dimensional structural characterization. Here, we demonstrate that cryo-electron microscopy can routinely resolve maps of RNA-only systems and that these maps enable subnanometer-resolution coordinate estimation when complemented with multidimensional chemical mapping and Rosetta DRRAFTER computational modeling. This hybrid 'Ribosolve' pipeline detects and falsifies homologies and conformational rearrangements in 11 previously unknown 119- to 338-nucleotide protein-free RNA structures: full-length Tetrahymena ribozyme, hc16 ligase with and without substrate, full-length Vibrio cholerae and Fusobacterium nucleatum glycine riboswitch aptamers with and without glycine, Mycobacterium SAM-IV riboswitch with and without S-adenosylmethionine, and the computer-designed ATP-TTR-3 aptamer with and without AMP. Simulation benchmarks, blind challenges, compensatory mutagenesis, cross-RNA homologies and internal controls demonstrate that Ribosolve can accurately resolve the global architectures of RNA molecules but does not resolve atomic details. These tests offer guidelines for making inferences in future RNA structural studies with similarly accelerated throughput.
- Biophysics Program, Stanford University, Stanford, CA, USA.
Organizational Affiliation: