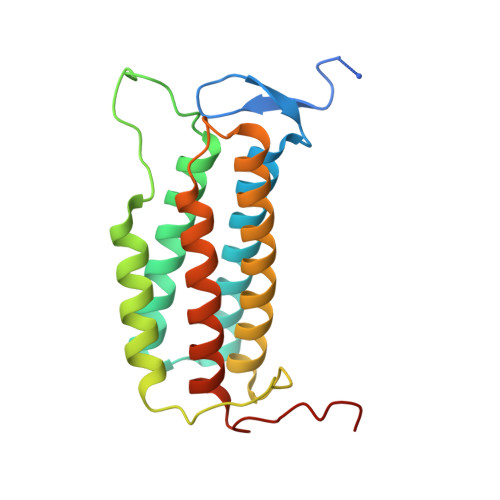
Mobile loop dynamics in adenosyltransferase control binding and reactivity of coenzyme B 12 .
Mascarenhas, R., Ruetz, M., McDevitt, L., Koutmos, M., Banerjee, R.(2020) Proc Natl Acad Sci U S A 117: 30412-30422
- PubMed: 33199623 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2007332117
- Primary Citation Related Structures:
6WGS, 6WGU, 6WGV, 6WH5 - PubMed Abstract:
Cobalamin is a complex organometallic cofactor that is processed and targeted via a network of chaperones to its dependent enzymes. AdoCbl (5'-deoxyadenosylcobalamin) is synthesized from cob(II)alamin in a reductive adenosylation reaction catalyzed by adenosyltransferase (ATR), which also serves as an escort, delivering AdoCbl to methylmalonyl-CoA mutase (MCM). The mechanism by which ATR signals that its cofactor cargo is ready (AdoCbl) or not [cob(II)alamin] for transfer to MCM, is not known. In this study, we have obtained crystallographic snapshots that reveal ligand-induced ordering of the N terminus of Mycobacterium tuberculosis ATR, which organizes a dynamic cobalamin binding site and exerts exquisite control over coordination geometry, reactivity, and solvent accessibility. Cob(II)alamin binds with its dimethylbenzimidazole tail splayed into a side pocket and its corrin ring buried. The cosubstrate, ATP, enforces a four-coordinate cob(II)alamin geometry, facilitating the unfavorable reduction to cob(I)alamin. The binding mode for AdoCbl is notably different from that of cob(II)alamin, with the dimethylbenzimidazole tail tucked under the corrin ring, displacing the N terminus of ATR, which is disordered. In this solvent-exposed conformation, AdoCbl undergoes facile transfer to MCM. The importance of the tail in cofactor handover from ATR to MCM is revealed by the failure of 5'-deoxyadenosylcobinamide, lacking the tail, to transfer. In the absence of MCM, ATR induces a sacrificial cobalt-carbon bond homolysis reaction in an unusual reversal of the heterolytic chemistry that was deployed to make the same bond. The data support an important role for the dimethylbenzimidazole tail in moving the cobalamin cofactor between active sites.
- Department of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109-0600.
Organizational Affiliation: