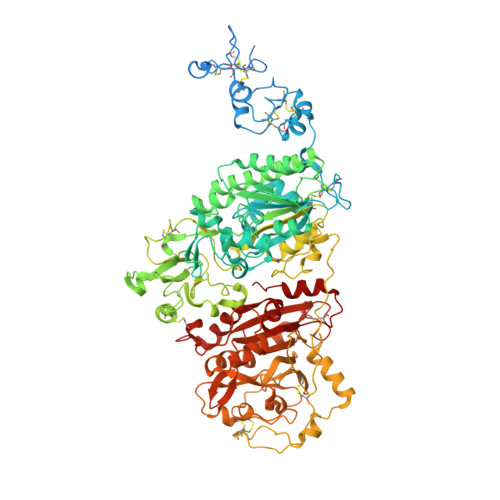
Crystal structures of human ENPP1 in apo and bound forms.
Dennis, M.L., Newman, J., Dolezal, O., Hattarki, M., Surjadi, R.N., Nuttall, S.D., Pham, T., Nebl, T., Camerino, M., Khoo, P.S., Monahan, B.J., Peat, T.S.(2020) Acta Crystallogr D Struct Biol 76: 889-898
- PubMed: 32876064 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798320010505
- Primary Citation Related Structures:
6WET, 6WEU, 6WEV, 6WEW, 6WFJ - PubMed Abstract:
Cancer is one of the leading causes of mortality in humans, and recent work has focused on the area of immuno-oncology, in which the immune system is used to specifically target cancerous cells. Ectonucleotide pyrophosphatase/phosphodiesterase 1 (ENPP1) is an emerging therapeutic target in human cancers owing to its role in degrading cyclic GMP-AMP (cGAMP), an agonist of the stimulator of interferon genes (STING). The available structures of ENPP1 are of the mouse enzyme, and no structures are available with anything other than native nucleotides. Here, the first X-ray crystal structures of the human ENPP1 enzyme in an apo form, with bound nucleotides and with two known inhibitors are presented. The availability of these structures and a robust crystallization system will allow the development of structure-based drug-design campaigns against this attractive cancer therapeutic target.
- Biomedical Manufacturing Program, CSIRO, 343 Royal Parade, Parkville, VIC 3052, Australia.
Organizational Affiliation: