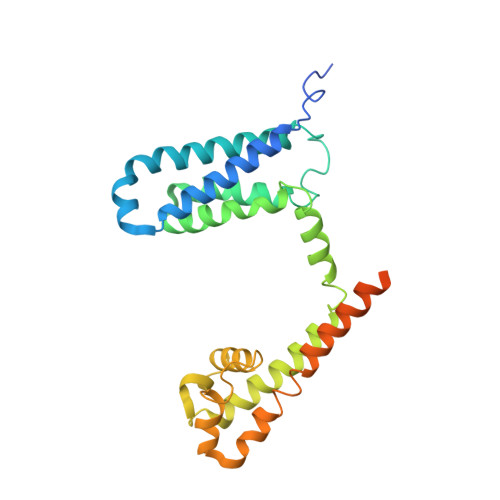
Employing NaChBac for cryo-EM analysis of toxin action on voltage-gated Na+channels in nanodisc.
Gao, S., Valinsky, W.C., On, N.C., Houlihan, P.R., Qu, Q., Liu, L., Pan, X., Clapham, D.E., Yan, N.(2020) Proc Natl Acad Sci U S A 117: 14187-14193
- PubMed: 32513729 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1922903117
- Primary Citation Related Structures:
6VWX, 6VX3, 6VXO, 6W6O - PubMed Abstract:
NaChBac, the first bacterial voltage-gated Na + (Na v ) channel to be characterized, has been the prokaryotic prototype for studying the structure-function relationship of Na v channels. Discovered nearly two decades ago, the structure of NaChBac has not been determined. Here we present the single particle electron cryomicroscopy (cryo-EM) analysis of NaChBac in both detergent micelles and nanodiscs. Under both conditions, the conformation of NaChBac is nearly identical to that of the potentially inactivated Na v Ab. Determining the structure of NaChBac in nanodiscs enabled us to examine gating modifier toxins (GMTs) of Na v channels in lipid bilayers. To study GMTs in mammalian Na v channels, we generated a chimera in which the extracellular fragment of the S3 and S4 segments in the second voltage-sensing domain from Na v 1.7 replaced the corresponding sequence in NaChBac. Cryo-EM structures of the nanodisc-embedded chimera alone and in complex with HuwenToxin IV (HWTX-IV) were determined to 3.5 and 3.2 Å resolutions, respectively. Compared to the structure of HWTX-IV-bound human Na v 1.7, which was obtained at an overall resolution of 3.2 Å, the local resolution of the toxin has been improved from ∼6 to ∼4 Å. This resolution enabled visualization of toxin docking. NaChBac can thus serve as a convenient surrogate for structural studies of the interactions between GMTs and Na v channels in a membrane environment.
- Department of Molecular Biology, Princeton University, Princeton, NJ 08544.
Organizational Affiliation: