Crystallographic and kinetic analyses of the FdsBG subcomplex of the cytosolic formate dehydrogenase FdsABG fromCupriavidus necator.
Young, T., Niks, D., Hakopian, S., Tam, T.K., Yu, X., Hille, R., Blaha, G.M.(2020) J Biological Chem 295: 6570-6585
- PubMed: 32249211
- DOI: https://doi.org/10.1074/jbc.RA120.013264
- Primary Citation Related Structures:
6VW7, 6VW8 - PubMed Abstract:
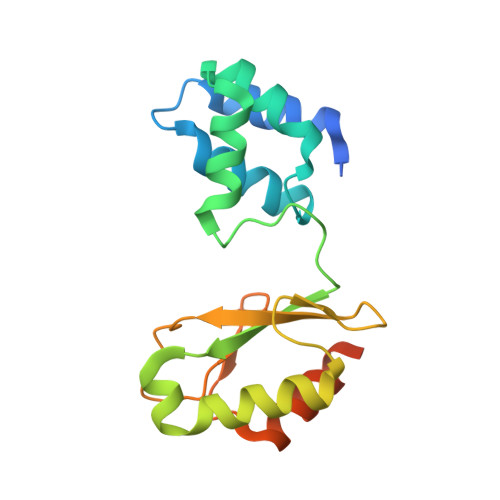
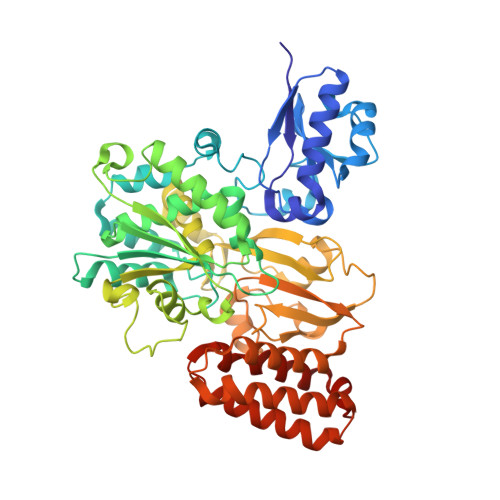
Formate oxidation to carbon dioxide is a key reaction in one-carbon compound metabolism, and its reverse reaction represents the first step in carbon assimilation in the acetogenic and methanogenic branches of many anaerobic organisms. The molybdenum-containing dehydrogenase FdsABG is a soluble NAD + -dependent formate dehydrogenase and a member of the NADH dehydrogenase superfamily. Here, we present the first structure of the FdsBG subcomplex of the cytosolic FdsABG formate dehydrogenase from the hydrogen-oxidizing bacterium Cupriavidus necator H16 both with and without bound NADH. The structures revealed that the two iron-sulfur clusters, Fe 4 S 4 in FdsB and Fe 2 S 2 in FdsG, are closer to the FMN than they are in other NADH dehydrogenases. Rapid kinetic studies and EPR measurements of rapid freeze-quenched samples of the NADH reduction of FdsBG identified a neutral flavin semiquinone, FMNH • , not previously observed to participate in NADH-mediated reduction of the FdsABG holoenzyme. We found that this semiquinone forms through the transfer of one electron from the fully reduced FMNH - , initially formed via NADH-mediated reduction, to the Fe 2 S 2 cluster. This Fe 2 S 2 cluster is not part of the on-path chain of iron-sulfur clusters connecting the FMN of FdsB with the active-site molybdenum center of FdsA. According to the NADH-bound structure, the nicotinamide ring stacks onto the re -face of the FMN. However, NADH binding significantly reduced the electron density for the isoalloxazine ring of FMN and induced a conformational change in residues of the FMN-binding pocket that display peptide-bond flipping upon NAD + binding in proper NADH dehydrogenases.
- Department of Biochemistry, University of California, Riverside, California 92521.
Organizational Affiliation: