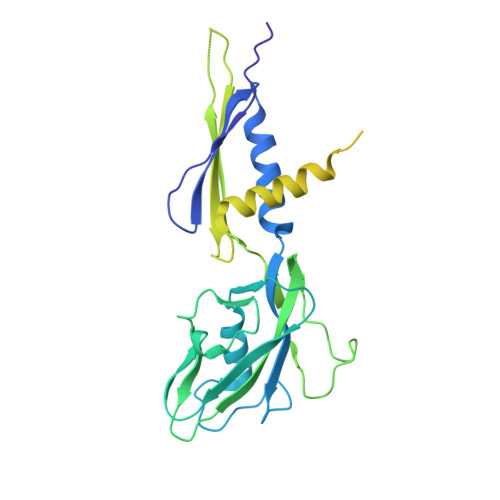
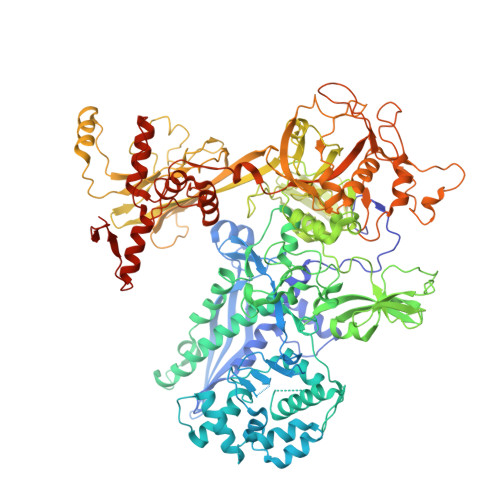
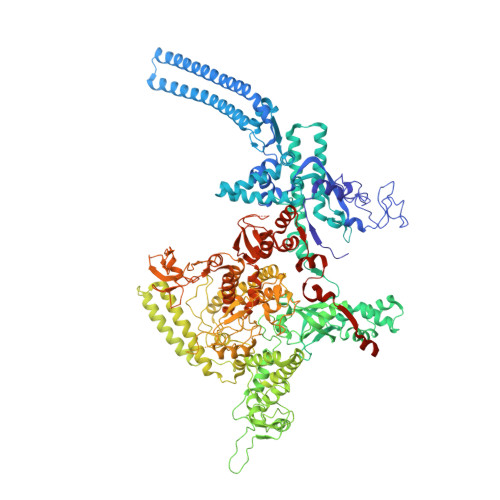
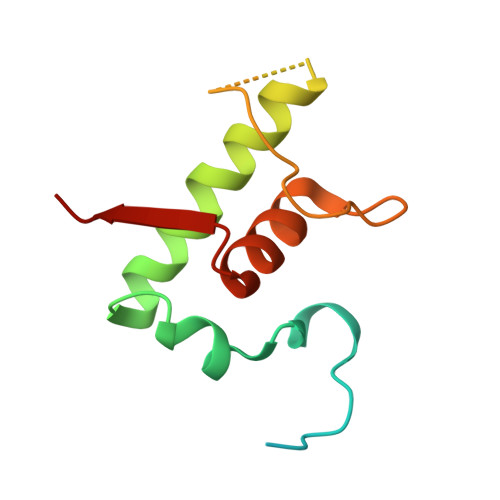
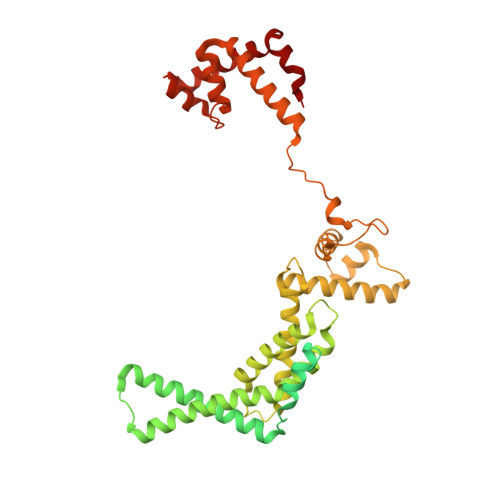
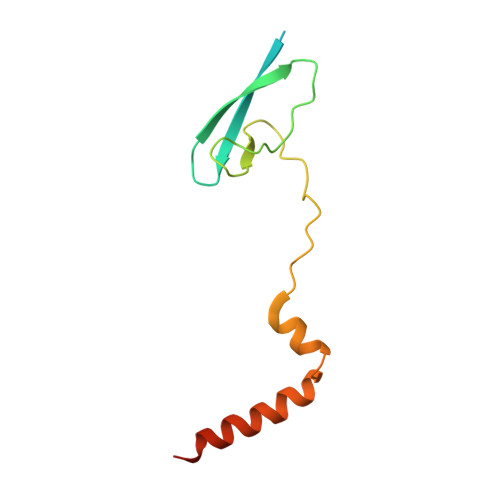
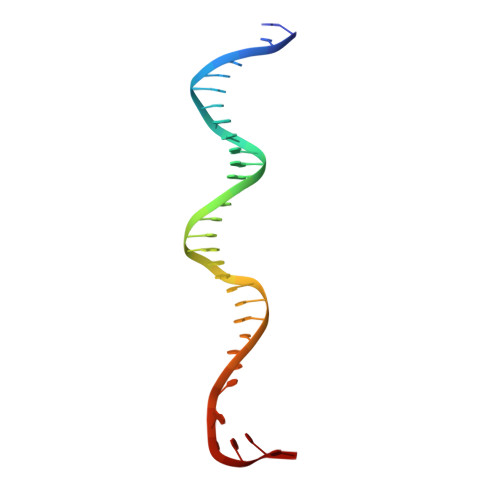
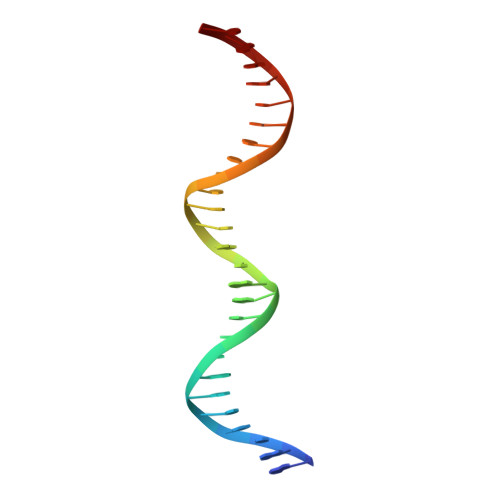
The antibiotic sorangicin A inhibits promoter DNA unwinding in a Mycobacterium tuberculosis rifampicin-resistant RNA polymerase.
Lilic, M., Chen, J., Boyaci, H., Braffman, N., Hubin, E.A., Herrmann, J., Muller, R., Mooney, R., Landick, R., Darst, S.A., Campbell, E.A.(2020) Proc Natl Acad Sci U S A 117: 30423-30432
- PubMed: 33199626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2013706117
- Primary Citation Related Structures:
6VVS, 6VVT, 6VVV, 6VVX, 6VVY, 6VVZ, 6VW0 - PubMed Abstract:
Rifampicin (Rif) is a first-line therapeutic used to treat the infectious disease tuberculosis (TB), which is caused by the pathogen Mycobacterium tuberculosis ( Mtb ). The emergence of Rif-resistant (Rif R ) Mtb presents a need for new antibiotics. Rif targets the enzyme RNA polymerase (RNAP). Sorangicin A (Sor) is an unrelated inhibitor that binds in the Rif-binding pocket of RNAP. Sor inhibits a subset of Rif R RNAPs, including the most prevalent clinical Rif R RNAP substitution found in Mtb infected patients (S456>L of the β subunit). Here, we present structural and biochemical data demonstrating that Sor inhibits the wild-type Mtb RNAP by a similar mechanism as Rif: by preventing the translocation of very short RNAs. By contrast, Sor inhibits the Rif R S456L enzyme at an earlier step, preventing the transition of a partially unwound promoter DNA intermediate to the fully opened DNA and blocking the template-strand DNA from reaching the active site in the RNAP catalytic center. By defining template-strand blocking as a mechanism for inhibition, we provide a mechanistic drug target in RNAP. Our finding that Sor inhibits the wild-type and mutant RNAPs through different mechanisms prompts future considerations for designing antibiotics against resistant targets. Also, we show that Sor has a better pharmacokinetic profile than Rif, making it a suitable starting molecule to design drugs to be used for the treatment of TB patients with comorbidities who require multiple medications.
- Laboratory of Molecular Biophysics, The Rockefeller University, New York, NY 10065.
Organizational Affiliation: