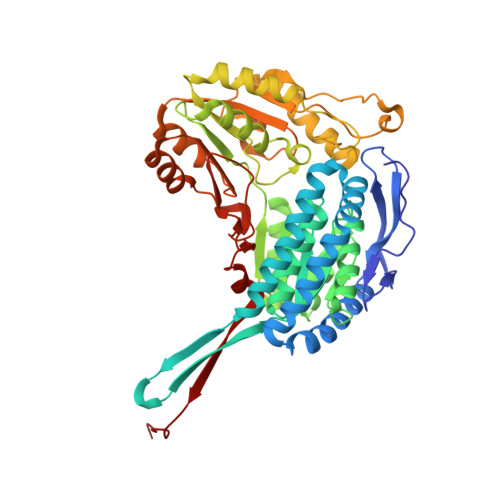
Inhibition, crystal structures, and in-solution oligomeric structure of aldehyde dehydrogenase 9A1.
Wyatt, J.W., Korasick, D.A., Qureshi, I.A., Campbell, A.C., Gates, K.S., Tanner, J.J.(2020) Arch Biochem Biophys 691: 108477-108477
- PubMed: 32717224 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.abb.2020.108477
- Primary Citation Related Structures:
6VR6, 6VWF - PubMed Abstract:
Aldehyde dehydrogenase 9A1 (ALDH9A1) is a human enzyme that catalyzes the NAD + -dependent oxidation of the carnitine precursor 4-trimethylaminobutyraldehyde to 4-N-trimethylaminobutyrate. Here we show that the broad-spectrum ALDH inhibitor diethylaminobenzaldehyde (DEAB) reversibly inhibits ALDH9A1 in a time-dependent manner. Possible mechanisms of inhibition include covalent reversible inactivation involving the thiohemiacetal intermediate and slow, tight-binding inhibition. Two crystal structures of ALDH9A1 are reported, including the first of the enzyme complexed with NAD + . One of the structures reveals the active conformation of the enzyme, in which the Rossmann dinucleotide-binding domain is fully ordered and the inter-domain linker adopts the canonical β-hairpin observed in other ALDH structures. The oligomeric structure of ALDH9A1 was investigated using analytical ultracentrifugation, small-angle X-ray scattering, and negative stain electron microscopy. These data show that ALDH9A1 forms the classic ALDH superfamily dimer-of-dimers tetramer in solution. Our results suggest that the presence of an aldehyde substrate and NAD + promotes isomerization of the enzyme into the active conformation.
- Department of Chemistry, University of Missouri, Columbia, MO, 65211, United States.
Organizational Affiliation: