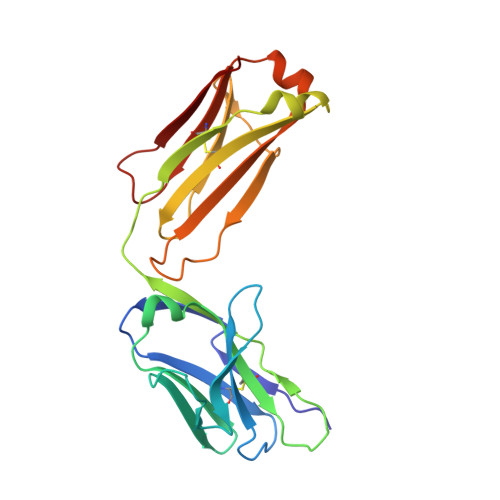
Recognition of an alpha-helical hairpin in P22 large terminase by a synthetic antibody fragment.
Lokareddy, R.K., Ko, Y.H., Hong, N., Doll, S.G., Paduch, M., Niederweis, M., Kossiakoff, A.A., Cingolani, G.(2020) Acta Crystallogr D Struct Biol 76: 876-888
- PubMed: 32876063 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798320009912
- Primary Citation Related Structures:
6VI1, 6VI2, 6XMI - PubMed Abstract:
The genome-packaging motor of tailed bacteriophages and herpesviruses is a multisubunit protein complex formed by several copies of a large (TerL) and a small (TerS) terminase subunit. The motor assembles transiently at the portal protein vertex of an empty precursor capsid to power the energy-dependent packaging of viral DNA. Both the ATPase and nuclease activities associated with genome packaging reside in TerL. Structural studies of TerL from bacteriophage P22 have been hindered by the conformational flexibility of this enzyme and its susceptibility to proteolysis. Here, an unbiased, synthetic phage-display Fab library was screened and a panel of high-affinity Fabs against P22 TerL were identified. This led to the discovery of a recombinant antibody fragment, Fab4, that binds a 33-amino-acid α-helical hairpin at the N-terminus of TerL with an equilibrium dissociation constant K d of 71.5 nM. A 1.51 Å resolution crystal structure of Fab4 bound to the TerL epitope (TLE) together with a 1.15 Å resolution crystal structure of the unliganded Fab4, which is the highest resolution ever achieved for a Fab, elucidate the principles governing the recognition of this novel helical epitope. TLE adopts two different conformations in the asymmetric unit and buries as much as 1250 Å 2 of solvent-accessible surface in Fab4. TLE recognition is primarily mediated by conformational changes in the third complementarity-determining region of the Fab4 heavy chain (CDR H3) that take place upon epitope binding. It is demonstrated that TLE can be introduced genetically at the N-terminus of a target protein, where it retains high-affinity binding to Fab4.
- Department of Biochemistry and Molecular Biology, Thomas Jefferson University, 1020 Locust Street, JAH-4E, Philadelphia, PA 19107, USA.
Organizational Affiliation: