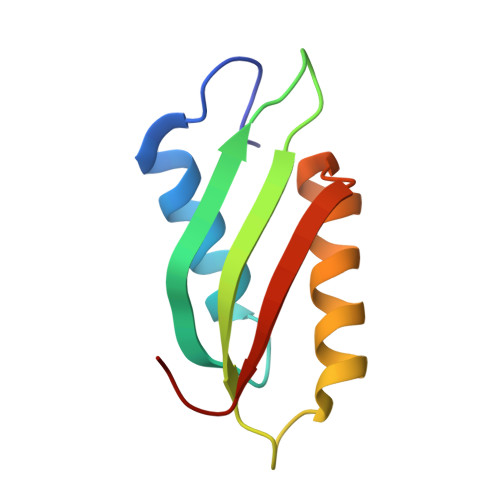
Structural and functional characterizations of the C-terminal domains of CzcD proteins.
Udagedara, S.R., La Porta, D.M., Spehar, C., Purohit, G., Hein, M.J.A., Fatmous, M.E., Casas Garcia, G.P., Ganio, K., McDevitt, C.A., Maher, M.J.(2020) J Inorg Biochem 208: 111087-111087
- PubMed: 32505855 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2020.111087
- Primary Citation Related Structures:
6VD8, 6VD9, 6VDA - PubMed Abstract:
Zinc is a potent antimicrobial component of the innate immune response at the host-pathogen interface. Bacteria subvert or resist host zinc insults by metal efflux pathways that include cation diffusion facilitator (CDF) proteins. The structural and functional examination of this protein class has been limited, with only the structures of the zinc transporter YiiP proteins from E. coli and Shewanella oneidensis described to date. Here, we determine the metal binding properties, solution quaternary structures and three dimensional architectures of the C-terminal domains of the metal transporter CzcD proteins from Cupriavidus metallidurans, Pseudomonas aeruginosa and Thermotoga maritima. We reveal significant diversity in the metal-binding properties and structures of these proteins and discover a potential novel mechanism for metal-promoted dimerization for the Cupriavidus metallidurans and Pseudomonas aeruginosa proteins.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne 3083, Australia.
Organizational Affiliation: