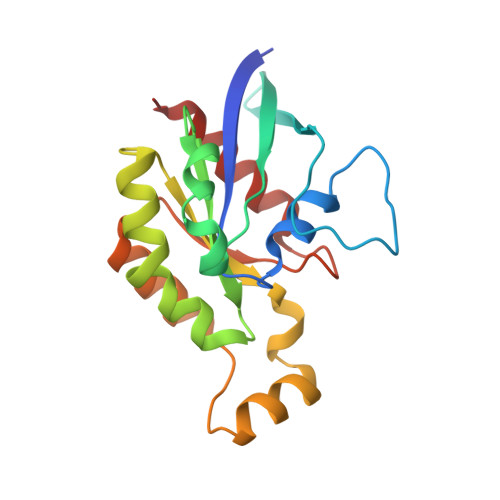
Structure of an inactive conformation of GTP-bound RhoA GTPase.
Lin, Y., Lu, S., Zhang, J., Zheng, Y.(2021) Structure 29: 553-563.e5
- PubMed: 33497604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.12.015
- Primary Citation Related Structures:
6V6M, 6V6U, 6V6V - PubMed Abstract:
By using 31 P NMR, we present evidence that the Rho family GTPase RhoA, similar to Ras GTPases, exists in an equilibrium of conformations when bound to GTP. High-resolution crystal structures of RhoA bound to the GTP analog GMPPNP and to GDP show that they display a similar overall inactive conformation. In contrast to the previously reported crystal structures of GTP analog-bound forms of two RhoA dominantly active mutants (G14V and Q63L), GMPPNP-bound RhoA assumes an open conformation in the Switch I loop with a previously unseen interaction between the γ-phosphate and Pro36, instead of the canonical Thr37. Molecular dynamics simulations found that the oncogenic RhoA G14V mutant displays a reduced flexibility in the Switch regions, consistent with a crystal structure of GDP-bound RhoA G14V . Thus, GDP- and GTP-bound RhoA can present similar inactive conformations, and the molecular dynamics in the Switch regions are likely to have a role in RhoA activation.
- Experimental Hematology and Cancer Biology, Cincinnati Children's Hospital Medical Center, University of Cincinnati College of Medicine, 3333 Burnet Avenue, Cincinnati, OH 45229, USA.
Organizational Affiliation: