Identification and Structure of a Multidonor Class of Head-Directed Influenza-Neutralizing Antibodies Reveal the Mechanism for Its Recurrent Elicitation.
Cheung, C.S., Fruehwirth, A., Paparoditis, P.C.G., Shen, C.H., Foglierini, M., Joyce, M.G., Leung, K., Piccoli, L., Rawi, R., Silacci-Fregni, C., Tsybovsky, Y., Verardi, R., Wang, L., Wang, S., Yang, E.S., Zhang, B., Zhang, Y., Chuang, G.Y., Corti, D., Mascola, J.R., Shapiro, L., Kwong, P.D., Lanzavecchia, A., Zhou, T.(2020) Cell Rep 32: 108088-108088
- PubMed: 32877670 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2020.108088
- Primary Citation Related Structures:
6URM - PubMed Abstract:
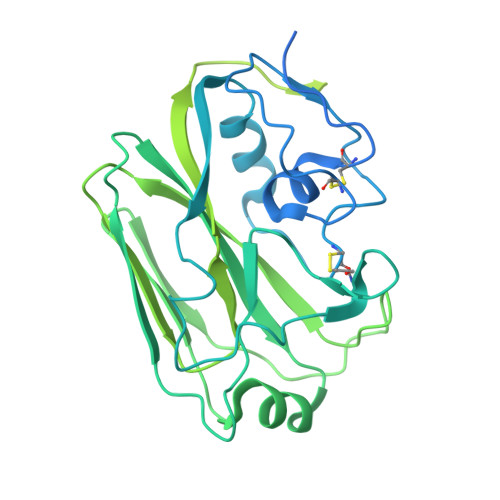
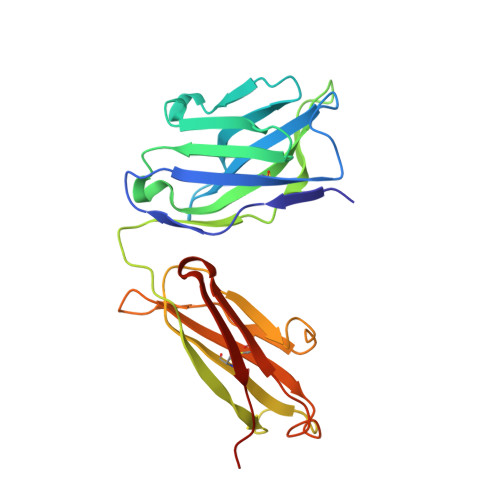
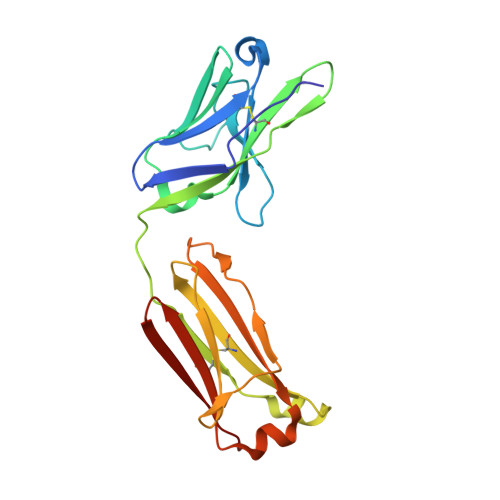
Multidonor antibodies are of interest for vaccine design because they can in principle be elicited in the general population by a common set of immunogens. For influenza, multidonor antibodies have been observed against the hemagglutinin (HA) stem, but not the immunodominant HA head. Here, we identify and characterize a multidonor antibody class (LPAF-a class) targeting the HA head. This class exhibits potent viral entry inhibition against H1N1 A/California/04/2009 (CA09) virus. LPAF-a class antibodies derive from the HV2-70 gene and contain a "Tyr-Gly-Asp"-motif, which occludes the HA-sialic acid binding site as revealed by a co-crystal structure with HA. Both germline-reverted and mature LPAF antibodies potently neutralize CA09 virus and have nanomolar affinities for CA09 HA. Moreover, increased frequencies for LPFA-a class antibodies are observed in humans after a single vaccination. Overall, this work highlights the identification of a multidonor class of head-directed influenza-neutralizing antibodies and delineates the mechanism of their recurrent elicitation in humans.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: