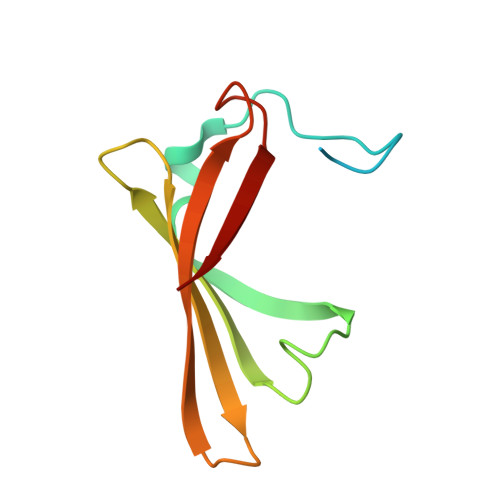
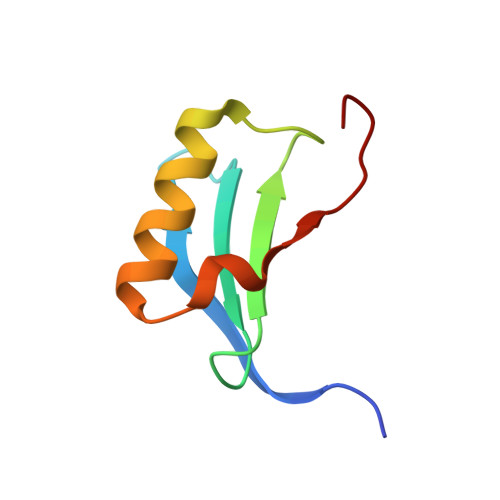
DiB-splits: nature-guided design of a novel fluorescent labeling split system.
Bozhanova, N.G., Gavrikov, A.S., Mishin, A.S., Meiler, J.(2020) Sci Rep 10: 11049-11049
- PubMed: 32632329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-67095-2
- Primary Citation Related Structures:
6UKK, 6UKL - PubMed Abstract:
Fluorogen-activating proteins (FAPs) are innovative fluorescent probes combining advantages of genetically-encoded proteins such as green fluorescent protein and externally added fluorogens that allow for highly tunable and on demand fluorescent signaling. Previously, a panel of green- and red-emitting FAPs has been created from bacterial lipocalin Blc (named DiBs). Here we present a rational design as well as functional and structural characterization of the first self-assembling FAP split system, DiB-splits. This new system decreases the size of the FAP label to ~8-12 kDa while preserving DiBs' unique properties: strong increase in fluorescence intensity of the chromophore upon binding, binding affinities to the chromophore in nanomolar to low micromolar range, and high photostability of the protein-ligand complex. These properties allow for use of DiB-splits for wide-field, confocal, and super-resolution fluorescence microscopy. DiB-splits also represent an attractive starting point for further design of a protein-protein interaction detection system as well as novel FAP-based sensors.
- Department of Chemistry, Center for Structural Biology, Vanderbilt University, Nashville, TN, 37235, USA.
Organizational Affiliation: