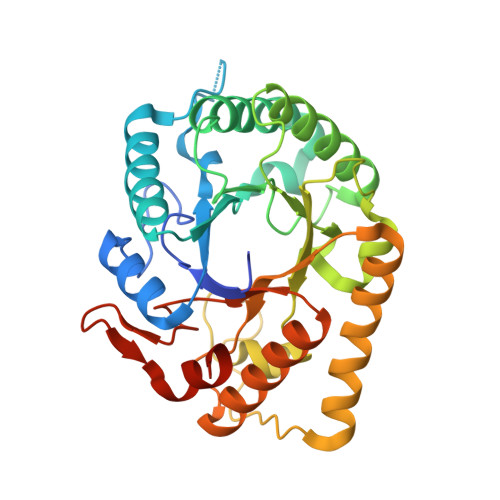
Identification of the Clostridial cellulose synthase and characterization of the cognate glycosyl hydrolase, CcsZ.
Scott, W., Lowrance, B., Anderson, A.C., Weadge, J.T.(2020) PLoS One 15: e0242686-e0242686
- PubMed: 33264329
- DOI: https://doi.org/10.1371/journal.pone.0242686
- Primary Citation Related Structures:
6UJE, 6UJF - PubMed Abstract:
Biofilms are community structures of bacteria enmeshed in a self-produced matrix of exopolysaccharides. The biofilm matrix serves numerous roles, including resilience and persistence, making biofilms a subject of research interest among persistent clinical pathogens of global health importance. Our current understanding of the underlying biochemical pathways responsible for biosynthesis of these exopolysaccharides is largely limited to Gram-negative bacteria. Clostridia are a class of Gram-positive, anaerobic and spore-forming bacteria and include the important human pathogens Clostridium perfringens, Clostridium botulinum and Clostridioides difficile, among numerous others. Several species of Clostridia have been reported to produce a biofilm matrix that contains an acetylated glucan linked to a series of hypothetical genes. Here, we propose a model for the function of these hypothetical genes, which, using homology modelling, we show plausibly encode a synthase complex responsible for polymerization, modification and export of an O-acetylated cellulose exopolysaccharide. Specifically, the cellulose synthase is homologous to that of the known exopolysaccharide synthases in Gram-negative bacteria. The remaining proteins represent a mosaic of evolutionary lineages that differ from the described Gram-negative cellulose exopolysaccharide synthases, but their predicted functions satisfy all criteria required for a functional cellulose synthase operon. Accordingly, we named these hypothetical genes ccsZABHI, for the Clostridial cellulose synthase (Ccs), in keeping with naming conventions for exopolysaccharide synthase subunits and to distinguish it from the Gram-negative Bcs locus with which it shares only a single one-to-one ortholog. To test our model and assess the identity of the exopolysaccharide, we subcloned the putative glycoside hydrolase encoded by ccsZ and solved the X-ray crystal structure of both apo- and product-bound CcsZ, which belongs to glycoside hydrolase family 5 (GH-5). Although not homologous to the Gram-negative cellulose synthase, which instead encodes the structurally distinct BcsZ belonging to GH-8, we show CcsZ displays specificity for cellulosic materials. This specificity of the synthase-associated glycosyl hydrolase validates our proposal that these hypothetical genes are responsible for biosynthesis of a cellulose exopolysaccharide. The data we present here allowed us to propose a model for Clostridial cellulose synthesis and serves as an entry point to an understanding of cellulose biofilm formation among class Clostridia.
- Department of Biology, Wilfrid Laurier University, Waterloo, ON, Canada.
Organizational Affiliation: