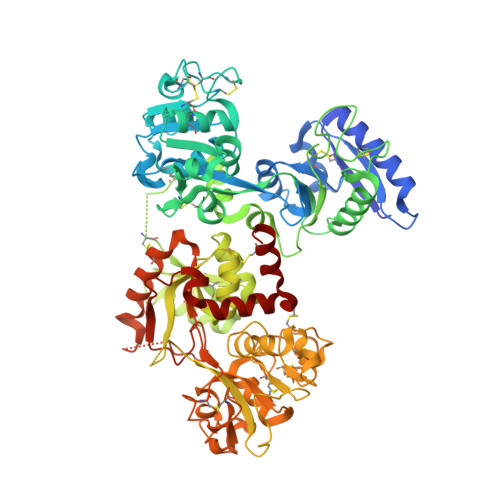
X-ray structure of chromium(III)-containing transferrin: First structure of a physiological Cr(III)-binding protein.
Petersen, C.M., Edwards, K.C., Gilbert, N.C., Vincent, J.B., Thompson, M.K.(2020) J Inorg Biochem 210: 111101-111101
- PubMed: 32650146 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2020.111101
- Primary Citation Related Structures:
6UJ6 - PubMed Abstract:
Transferrin, the Fe(III) transport protein in mammalian blood, has been suggested to also serve as a Cr(III) transporter and as part of a Cr(III) detoxification system; however, the structure of the metal-binding sites has never been fully elucidated with bound Cr(III). Chromium(III)-transferrin was crystallized in the presence of the synergistic anion malonate. In the crystals, the protein exists with a closed C-terminal lobe containing a Cr(III) ion and an open, unoccupied N-terminal lobe. The overall structure and the metal ion environments are extremely similar to those of Fe(III)- and Ti(IV)-containing transferrin crystallized under comparable conditions. The octahedral coordination about the Cr(III) is comprised of four ligands provided by the protein (two tyrosine residues, a histidine residue, and an aspartate residue) and a chelating malonate anion. This represents the first crystal structure of a Cr(III)-containing protein that binds Cr(III) as part of its physiological function.
- Department of Chemistry & Biochemistry, The University of Alabama, 250 Hackberry Lane, Tuscaloosa, AL 35487-0336, USA.
Organizational Affiliation: