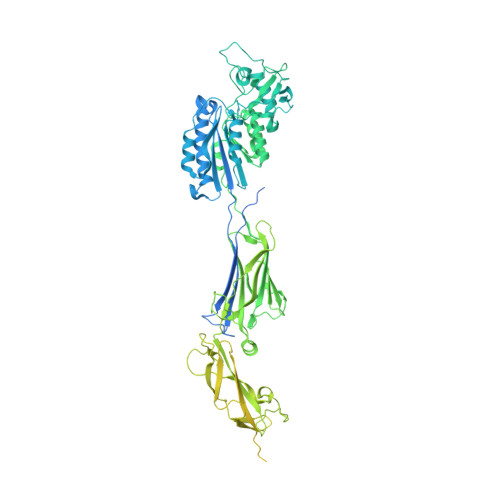
Porphyromonas gingivalis fimbrial protein Mfa5 contains a von Willebrand factor domain and an intramolecular isopeptide.
Heidler, T.V., Ernits, K., Ziolkowska, A., Claesson, R., Persson, K.(2021) Commun Biol 4: 106-106
- PubMed: 33495563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-01621-w
- Primary Citation Related Structures:
6TNJ, 6TO1 - PubMed Abstract:
The Gram-negative bacterium Porphyromonas gingivalis is a secondary colonizer of the oral biofilm and is involved in the onset and progression of periodontitis. Its fimbriae, of type-V, are important for attachment to other microorganisms in the biofilm and for adhesion to host cells. The fimbriae are assembled from five proteins encoded by the mfa1 operon, of which Mfa5 is one of the ancillary tip proteins. Here we report the X-ray structure of the N-terminal half of Mfa5, which reveals a von Willebrand factor domain and two IgG-like domains. One of the IgG-like domains is stabilized by an intramolecular isopeptide bond, which is the first such bond observed in a Gram-negative bacterium. These features make Mfa5 structurally more related to streptococcal adhesins than to the other P. gingivalis Mfa proteins. The structure reported here indicates that horizontal gene transfer has occurred among the bacteria within the oral biofilm.
- Department of Chemistry, Umeå Centre for Microbial Research (UCMR), Umeå University, 90187, Umeå, Sweden.
Organizational Affiliation: