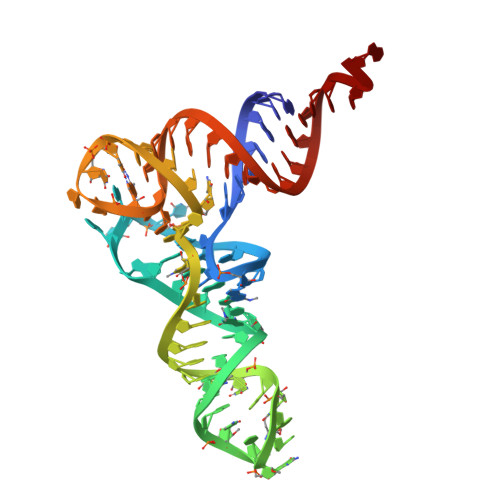
Crystal structure of yeast phenylalanine transfer RNA. I. Crystallographic refinement.
Sussman, J.L., Holbrook, S.R., Warrant, R.W., Church, G.M., Kim, S.H.(1978) J Mol Biology 123: 607-630
- PubMed: 357742 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(78)90209-7
- Primary Citation Related Structures:
6TNA