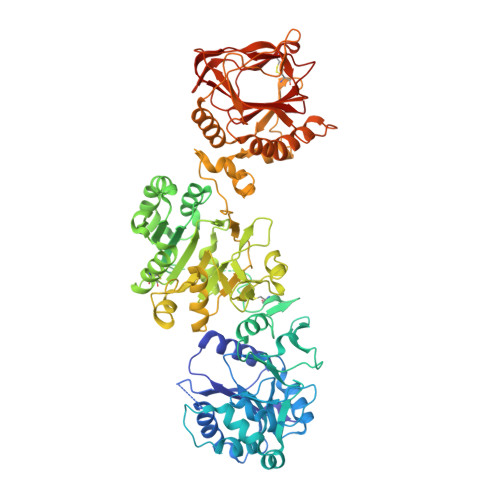
Identification of Regulatory Molecular 'Hot Spots' for LH/PLOD Collagen Glycosyltransferase Activity
Mattoteia, D., Chiapparino, A., Fumagalli, M., De Marco, M., De Giorgi, F., Negro, L., Pinnola, A., Faravelli, S., Roscioli, T., Scietti, L., Forneris, F.(2023) Int J Mol Sci