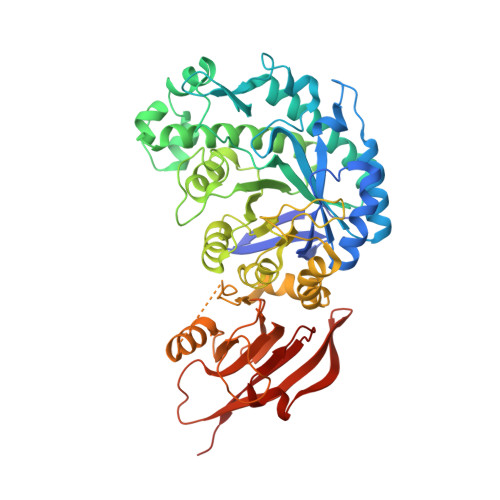
Mechanistic Insights into the Chaperoning of Human Lysosomal-Galactosidase Activity: Highly Functionalized Aminocyclopentanes and C -5a-Substituted Derivatives of 4- epi -Isofagomine.
Weber, P., Thonhofer, M., Averill, S., Davies, G.J., Santana, A.G., Muller, P., Nasseri, S.A., Offen, W.A., Pabst, B.M., Paschke, E., Schalli, M., Torvisco, A., Tschernutter, M., Tysoe, C., Windischhofer, W., Withers, S.G., Wolfsgruber, A., Wrodnigg, T.M., Stutz, A.E.(2020) Molecules 25
- PubMed: 32899288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules25174025
- Primary Citation Related Structures:
6TBF, 6TBG, 6TBH, 6TBI, 6TBJ, 6TBK - PubMed Abstract:
Glycosidase inhibitors have shown great potential as pharmacological chaperones for lysosomal storage diseases. In light of this, a series of new cyclopentanoid β-galactosidase inhibitors were prepared and their inhibitory and pharmacological chaperoning activities determined and compared with those of lipophilic analogs of the potent β-d-galactosidase inhibitor 4- epi -isofagomine. Structure-activity relationships were investigated by X-ray crystallography as well as by alterations in the cyclopentane moiety such as deoxygenation and replacement by fluorine of a "strategic" hydroxyl group. New compounds have revealed highly promising activities with a range of β-galactosidase-compromised human cell lines and may serve as leads towards new pharmacological chaperones for G M1 -gangliosidosis and Morquio B disease.
- Glycogroup, Institute of Chemistry and Technology of Biobased Systems, Graz University of Technology, Stremayrgasse 9, A-8010 Graz, Austria.
Organizational Affiliation: