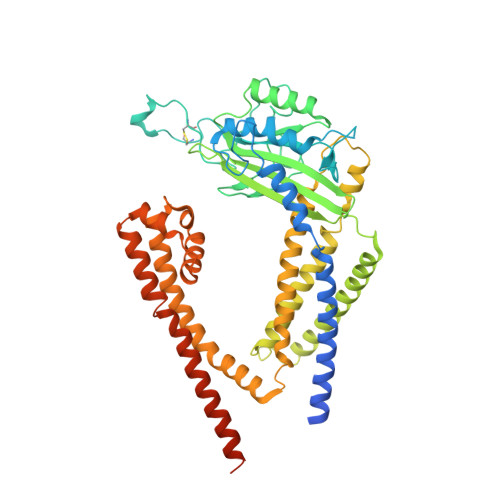
Lipid Interactions of a Ciliary Membrane TRP Channel: Simulation and Structural Studies of Polycystin-2.
Wang, Q., Corey, R.A., Hedger, G., Aryal, P., Grieben, M., Nasrallah, C., Baronina, A., Pike, A.C.W., Shi, J., Carpenter, E.P., Sansom, M.S.P.(2020) Structure 28: 169-184.e5
- PubMed: 31806353 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.11.005
- Primary Citation Related Structures:
6T9N, 6T9O - PubMed Abstract:
Polycystin-2 (PC2) is a transient receptor potential (TRP) channel present in ciliary membranes of the kidney. PC2 shares a transmembrane fold with other TRP channels, in addition to an extracellular domain found in TRPP and TRPML channels. Using molecular dynamics (MD) simulations and cryoelectron microscopy we identify and characterize PIP 2 and cholesterol interactions with PC2. PC2 is revealed to have a PIP binding site close to the equivalent vanilloid/lipid binding site in the TRPV1 channel. A 3.0-Å structure reveals a binding site for cholesterol on PC2. Cholesterol interactions with the channel at this site are characterized by MD simulations. The two classes of lipid binding sites are compared with sites observed in other TRPs and in Kv channels. These findings suggest PC2, in common with other ion channels, may be modulated by both PIPs and cholesterol, and position PC2 within an emerging model of the roles of lipids in the regulation and organization of ciliary membranes.
- Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, UK; Structural Genomics Consortium, University of Oxford, Old Road Campus Research Building, Roosevelt Drive, Oxford OX3 7DQ, UK.
Organizational Affiliation: