FtsK in motion reveals its mechanism for double-stranded DNA translocation.
Jean, N.L., Rutherford, T.J., Lowe, J.(2020) Proc Natl Acad Sci U S A 117: 14202-14208
- PubMed: 32513722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2001324117
- Primary Citation Related Structures:
6T8B, 6T8G, 6T8O - PubMed Abstract:
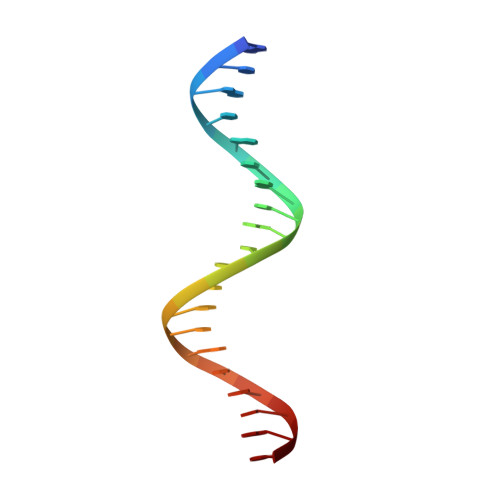
FtsK protein contains a fast DNA motor that is involved in bacterial chromosome dimer resolution. During cell division, FtsK translocates double-stranded DNA until both dif recombination sites are placed at mid cell for subsequent dimer resolution. Here, we solved the 3.6-Å resolution electron cryo-microscopy structure of the motor domain of FtsK while translocating on its DNA substrate. Each subunit of the homo-hexameric ring adopts a unique conformation and one of three nucleotide states. Two DNA-binding loops within four subunits form a pair of spiral staircases within the ring, interacting with the two DNA strands. This suggests that simultaneous conformational changes in all ATPase domains at each catalytic step generate movement through a mechanism related to filament treadmilling. While the ring is only rotating around the DNA slowly, it is instead the conformational states that rotate around the ring as the DNA substrate is pushed through.
- Structural Studies Division, Medical Research Council Laboratory of Molecular Biology, Cambridge CB2 0QH, United Kingdom.
Organizational Affiliation: