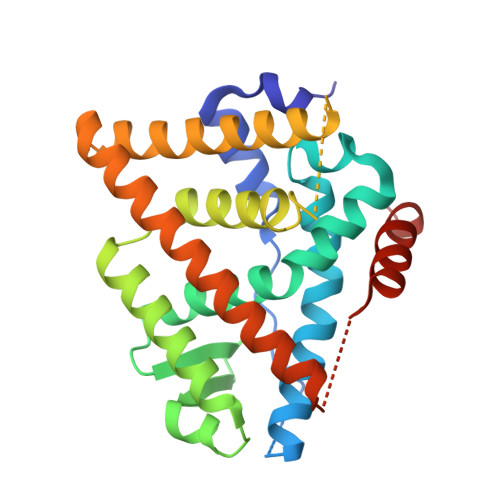
Building Bridges in a Series of Estrogen Receptor Degraders: An Application of Metathesis in Medicinal Chemistry.
Scott, J.S., Breed, J., Carbajo, R.J., Davey, P.R., Greenwood, R., Huynh, H.K., Klinowska, T., Morrow, C.J., Moss, T.A., Polanski, R., Nissink, J.W.M., Varnes, J., Yang, B.(2019) ACS Med Chem Lett 10: 1492-1497
- PubMed: 31620239 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.9b00370
- Primary Citation Related Structures:
6SQ0, 6SUO - PubMed Abstract:
Herein we report the use of metathesis to construct a novel tetracyclic core in a series of estrogen receptor degraders. This improved the chemical stability, as assessed using an NMR-MS based assay, and gave a molecule with excellent physicochemical properties and pharmacokinetics in rat. X-ray crystallography established minimal perturbation of the bridged compounds relative to the unbridged analogues in the receptor binding pocket. Unfortunately, despite retaining excellent binding to ERα, this adversely affected the ability of the compounds to degrade the receptor.
- Oncology R&D, AstraZeneca, Cambridge CB4 0WG, United Kingdom.
Organizational Affiliation: