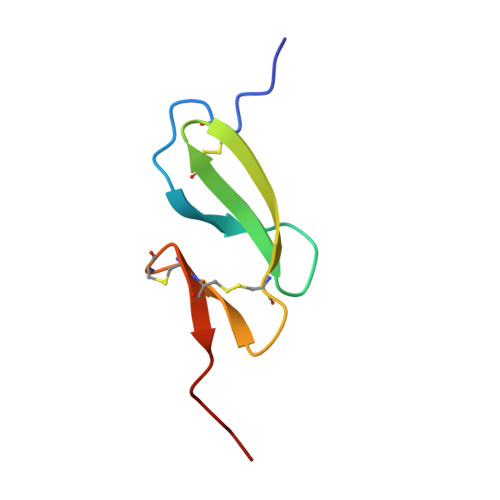
NMR and Fluorescence Spectroscopies Reveal the Preorganized Binding Site in Family 14 Carbohydrate-Binding Module from Human Chitotriosidase.
Madland, E., Crasson, O., Vandevenne, M., Sorlie, M., Aachmann, F.L.(2019) ACS Omega 4: 21975-21984
- PubMed: 31891077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.9b03043
- Primary Citation Related Structures:
6SO0 - PubMed Abstract:
Carbohydrate-binding modules (CBM) play important roles in targeting and increasing the concentration of carbohydrate active enzymes on their substrates. Using NMR to get the solution structure of CBM14, we can gain insight into secondary structure elements and intramolecular interactions with our assigned nuclear overhauser effect peaks. This reveals that two conserved aromatic residues (Phe437 and Phe456) make up the hydrophobic core of the CBM. These residues are also responsible for connecting the two β-sheets together, by being part of β2 and β4, respectively, and together with disulfide bridges, they create CBM14's characteristic "hevein-like" fold. Most CBMs rely on aromatic residues for substrate binding; however, CBM14 contains just a single tryptophan (Trp465) that together with Asn466 enables substrate binding. Interestingly, an alanine mutation of a single residue (Leu454) located behind Trp465 renders the CBM incapable of binding. Fluorescence spectroscopy performed on this mutant reveals a significant blue shift, as well as a minor blue shift for its neighbor Val455. The reduction in steric hindrance causes the tryptophan to be buried into the hydrophobic core of the structure and therefore suggests a preorganized binding site for this CBM. Our results show that both Trp465 and Asn466 are affected when CBM14 interacts with both (GlcNAc) 3 and β-chitin, that the binding interactions are weak, and that CBM14 displays a slightly higher affinity toward β-chitin.
- Department of Biotechnology and Food Science, Norwegian Biopolymer Laboratory (NOBIPOL), NTNU Norwegian University of Science and Technology, Trondheim 7491, Norway.
Organizational Affiliation: