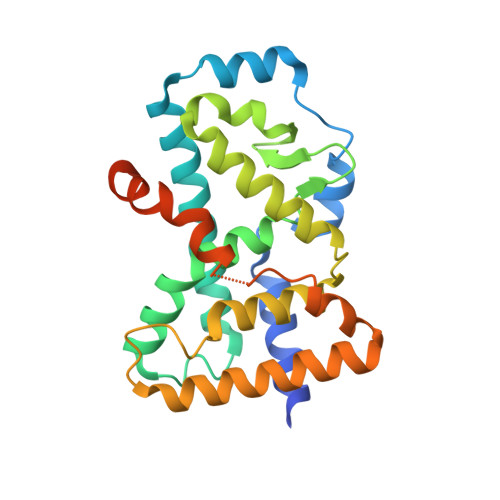
Crystal structure of human ROR gamma LBD in complex with a (quinolinoxymethyl)benzamide inverse agonist
Amaudrut, J., Argiriadi, M.A., Barth, M., Breinlinger, E.C., Calderwood, D.J., Cusack, K.P., Kort, M.E., Montalbetti, C., Potin, D., Poupardin, O., Spitzer, L.To be published.