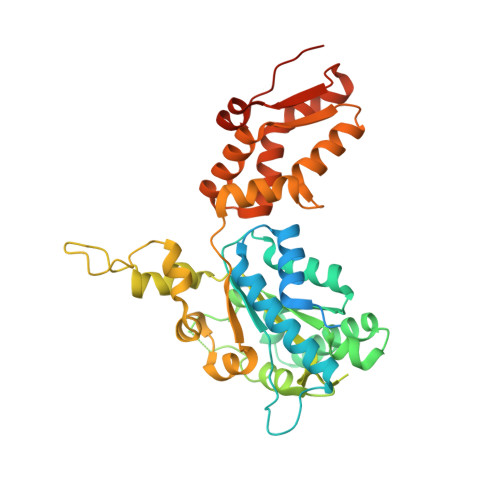
Cryo-EM structure of the ClpXP protein degradation machinery.
Gatsogiannis, C., Balogh, D., Merino, F., Sieber, S.A., Raunser, S.(2019) Nat Struct Mol Biol 26: 946-954
- PubMed: 31582852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0304-0
- Primary Citation Related Structures:
6SFW, 6SFX - PubMed Abstract:
The ClpXP machinery is a two-component protease complex that performs targeted protein degradation in bacteria and mitochondria. The complex consists of the AAA+ chaperone ClpX and the peptidase ClpP. The hexameric ClpX utilizes the energy of ATP binding and hydrolysis to engage, unfold and translocate substrates into the catalytic chamber of tetradecameric ClpP, where they are degraded. Formation of the complex involves a symmetry mismatch, because hexameric AAA+ rings bind axially to the opposing stacked heptameric rings of the tetradecameric ClpP. Here we present the cryo-EM structure of ClpXP from Listeria monocytogenes. We unravel the heptamer-hexamer binding interface and provide novel insight into the ClpX-ClpP cross-talk and activation mechanism. Comparison with available crystal structures of ClpP and ClpX in different states allows us to understand important aspects of the complex mode of action of ClpXP and provides a structural framework for future pharmacological applications.
- Department of Structural Biochemistry, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: