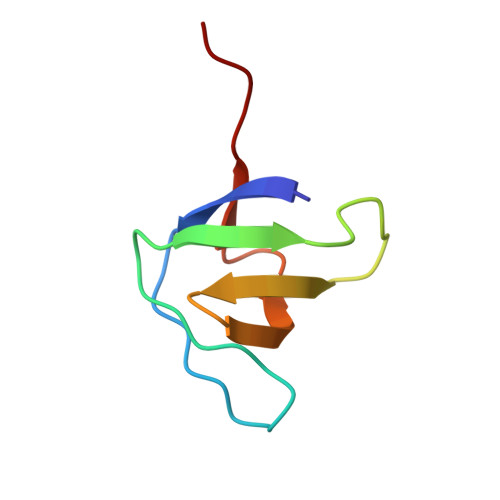
Crystal structure of the SH3 domain of growth factor receptor-bound protein 2.
Bolgov, A., Korban, S., Luzik, D., Zhemkov, V., Kim, M., Rogacheva, O., Bezprozvanny, I.(2020) Acta Crystallogr F Struct Biol Commun 76: 263-270
- PubMed: 32510467 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20007232
- Primary Citation Related Structures:
6SDF - PubMed Abstract:
This study presents the crystal structure of the N-terminal SH3 (SH3N) domain of growth factor receptor-bound protein 2 (Grb2) at 2.5 Å resolution. Grb2 is a small (215-amino-acid) adaptor protein that is widely expressed and involved in signal transduction/cell communication. The crystal structure of full-length Grb2 has previously been reported (PDB entry 1gri). The structure of the isolated SH3N domain is consistent with the full-length structure. The structure of the isolated SH3N domain was solved at a higher resolution (2.5 Å compared with 3.1 Å for the previously deposited structure) and made it possible to resolve some of the loops that were missing in the full-length structure. In addition, interactions between the carboxy-terminal region of the SH3N domain and the Sos1-binding sites were observed in the structure of the isolated domain. Analysis of these interactions provided new information about the ligand-binding properties of the SH3N domain of Grb2.
- Laboratory of Molecular Neurodegeneration, Institute of Biomedical systems and Biotechnologies, Peter the Great St Petersburg Polytechnic University, St Petersburg 194021, Russian Federation.
Organizational Affiliation: