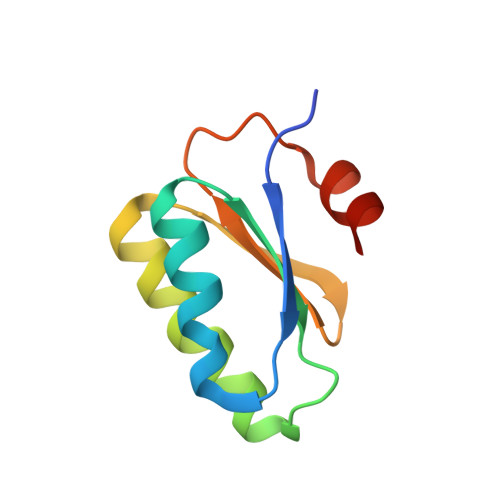
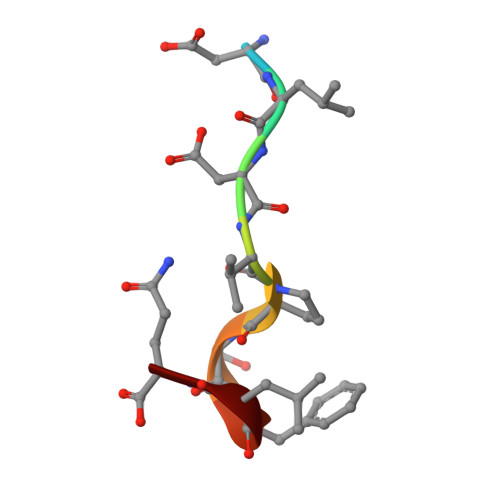
Essential dynamic interdependence of FtsZ and SepF for Z-ring and septum formation in Corynebacterium glutamicum.
Sogues, A., Martinez, M., Gaday, Q., Ben Assaya, M., Grana, M., Voegele, A., VanNieuwenhze, M., England, P., Haouz, A., Chenal, A., Trepout, S., Duran, R., Wehenkel, A.M., Alzari, P.M.(2020) Nat Commun 11: 1641-1641
- PubMed: 32242019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-15490-8
- Primary Citation Related Structures:
6SAT, 6SCP, 6SCQ, 6SCS - PubMed Abstract:
The mechanisms of Z-ring assembly and regulation in bacteria are poorly understood, particularly in non-model organisms. Actinobacteria, a large bacterial phylum that includes the pathogen Mycobacterium tuberculosis, lack the canonical FtsZ-membrane anchors and Z-ring regulators described for E. coli. Here we investigate the physiological function of Corynebacterium glutamicum SepF, the only cell division-associated protein from Actinobacteria known to interact with the conserved C-terminal tail of FtsZ. We show an essential interdependence of FtsZ and SepF for formation of a functional Z-ring in C. glutamicum. The crystal structure of the SepF-FtsZ complex reveals a hydrophobic FtsZ-binding pocket, which defines the SepF homodimer as the functional unit, and suggests a reversible oligomerization interface. FtsZ filaments and lipid membranes have opposing effects on SepF polymerization, indicating that SepF has multiple roles at the cell division site, involving FtsZ bundling, Z-ring tethering and membrane reshaping activities that are needed for proper Z-ring assembly and function.
- Unité de Microbiologie Structurale, Institut Pasteur, CNRS UMR 3528, Université de Paris, 75015, Paris, France.
Organizational Affiliation: