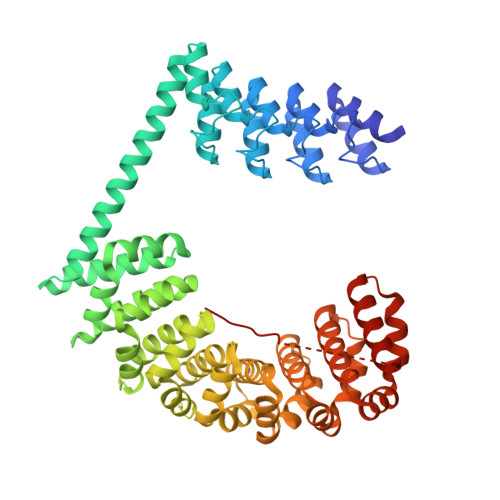
Rigid fusions of designed helical repeat binding proteins efficiently protect a binding surface from crystal contacts.
Ernst, P., Honegger, A., van der Valk, F., Ewald, C., Mittl, P.R.E., Pluckthun, A.(2019) Sci Rep 9: 16162-16162
- PubMed: 31700118
- DOI: https://doi.org/10.1038/s41598-019-52121-9
- Primary Citation of Related Structures:
6SA6, 6SA7, 6SA8 - PubMed Abstract:
Designed armadillo repeat proteins (dArmRPs) bind extended peptides in a modular way. The consensus version recognises alternating arginines and lysines, with one dipeptide per repeat. For generating new binding specificities, the rapid and robust analysis by crystallography is key. Yet, we have previously found that crystal contacts can strongly influence this analysis, by displacing the peptide and potentially distorting the overall geometry of the scaffold. Therefore, we now used protein design to minimise these effects and expand the previously described concept of shared helices to rigidly connect dArmRPs and designed ankyrin repeat proteins (DARPins), which serve as a crystallisation chaperone. To shield the peptide-binding surface from crystal contacts, we rigidly fused two DARPins to the N- and C-terminal repeat of the dArmRP and linked the two DARPins by a disulfide bond. In this ring-like structure, peptide binding, on the inside of the ring, is very regular and undistorted, highlighting the truly modular binding mode. Thus, protein design was utilised to construct a well crystallising scaffold that prevents interference from crystal contacts with peptide binding and maintains the equilibrium structure of the dArmRP. Rigid DARPin-dArmRPs fusions will also be useful when chimeric binding proteins with predefined geometries are required.
- Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, 8057, Zürich, Switzerland.
Organizational Affiliation: