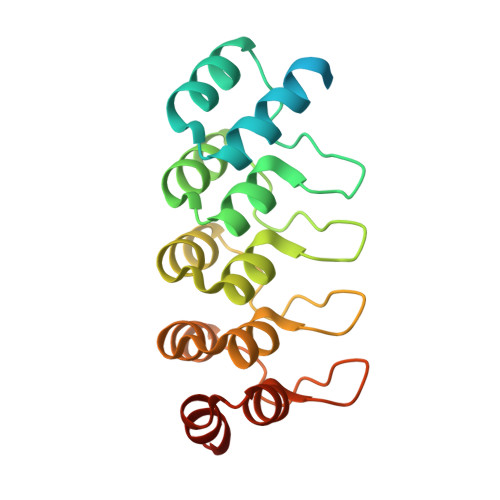
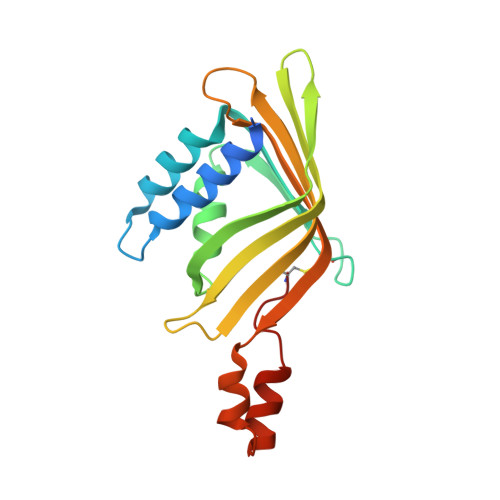
Rotational symmetry of the structured Chip/LDB-SSDP core module of the Wnt enhanceosome.
Renko, M., Fiedler, M., Rutherford, T.J., Schaefer, J.V., Pluckthun, A., Bienz, M.(2019) Proc Natl Acad Sci U S A 116: 20977-20983
- PubMed: 31570581
- DOI: https://doi.org/10.1073/pnas.1912705116
- Primary Citation of Related Structures:
6S9R, 6S9S, 6S9T - PubMed Abstract:
The Chip/LIM-domain binding protein (LDB)-single-stranded DNA-binding protein (SSDP) (ChiLS) complex controls numerous cell-fate decisions in animal cells, by mediating transcription of developmental control genes via remote enhancers. ChiLS is recruited to these enhancers by lineage-specific LIM-domain proteins that bind to its Chip/LDB subunit. ChiLS recently emerged as the core module of the Wnt enhanceosome, a multiprotein complex that primes developmental control genes for timely Wnt responses. ChiLS binds to NPFxD motifs within Pygopus (Pygo) and the Osa/ARID1A subunit of the BAF chromatin remodeling complex, which could synergize with LIM proteins in tethering ChiLS to enhancers. Chip/LDB and SSDP both contain N-terminal dimerization domains that constitute the bulk of their structured cores. Here, we report the crystal structures of these dimerization domains, in part aided by DARPin chaperones. We conducted systematic surface scanning by structure-designed mutations, followed by in vitro and in vivo binding assays, to determine conserved surface residues required for binding between Chip/LDB, SSDP, and Pygo-NPFxD. Based on this, and on the 4:2 (SSDP-Chip/LDB) stoichiometry of ChiLS, we derive a highly constrained structural model for this complex, which adopts a rotationally symmetrical SSDP 2 -LDB 2 -SSDP 2 architecture. Integrity of ChiLS is essential for Pygo binding, and our mutational analysis places the NPFxD pockets on either side of the Chip/LDB dimer, each flanked by an SSDP dimer. The symmetry and multivalency of ChiLS underpin its function as an enhancer module integrating Wnt signals with lineage-specific factors to operate context-dependent transcriptional switches that are pivotal for normal development and cancer.
- Medical Research Council Laboratory of Molecular Biology, CB2 0QH Cambridge, United Kingdom.
Organizational Affiliation: