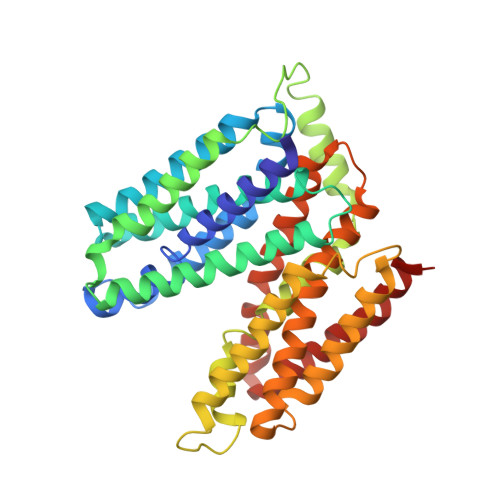
Structure of a proton-dependent lipid transporter involved in lipoteichoic acids biosynthesis.
Zhang, B., Liu, X., Lambert, E., Mas, G., Hiller, S., Veening, J.W., Perez, C.(2020) Nat Struct Mol Biol 27: 561-569
- PubMed: 32367070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0425-5
- Primary Citation Related Structures:
6S7V - PubMed Abstract:
Lipoteichoic acids (LTAs) are essential cell-wall components in Gram-positive bacteria, including the human pathogen Staphylococcus aureus, contributing to cell adhesion, cell division and antibiotic resistance. Genetic evidence has suggested that LtaA is the flippase that mediates the translocation of the lipid-linked disaccharide that anchors LTA to the cell membrane, a rate-limiting step in S. aureus LTA biogenesis. Here, we present the structure of LtaA, describe its flipping mechanism and show its functional relevance for S. aureus fitness. We demonstrate that LtaA is a proton-coupled antiporter flippase that contributes to S. aureus survival under physiological acidic conditions. Our results provide foundations for the development of new strategies to counteract S. aureus infections.
- Biozentrum, University of Basel, Basel, Switzerland.
Organizational Affiliation: