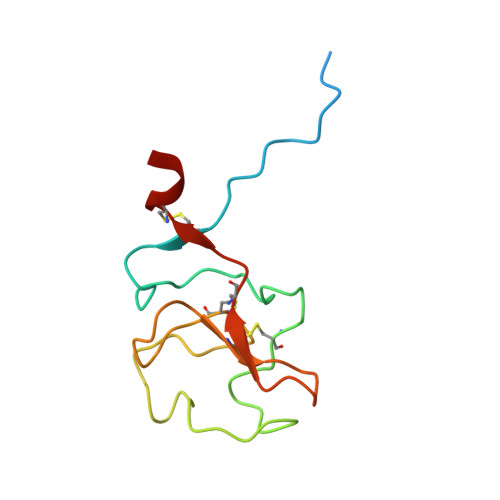
High resolution structure of human apolipoprotein (a) kringle IV type 2: beyond the lysine binding site.
Santonastaso, A., Maggi, M., De Jonge, H., Scotti, C.(2020) J Lipid Res 61: 1687-1696
- PubMed: 32907988 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1194/jlr.RA120001023
- Primary Citation Related Structures:
6RX7 - PubMed Abstract:
Lipoprotein (a) [Lp(a)] is characterized by an LDL-like composition in terms of lipids and apoB100, and by one copy of a unique glycoprotein, apo(a). The apo(a) structure is mainly based on the repetition of tandem kringle domains with high homology to plasminogen kringles 4 and 5. Among them, kringle IV type 2 (KIV-2) is present in a highly variable number of genetically encoded repeats, whose length is inversely related to Lp(a) plasma concentration and cardiovascular risk. Despite it being the major component of apo(a), the actual function of KIV-2 is still unclear. Here, we describe the first high-resolution crystallographic structure of this domain. It shows a general fold very similar to other KIV domains with high and intermediate affinity for the lysine analog, ε-aminocaproic acid. Interestingly, KIV-2 presents a lysine binding site (LBS) with a unique shape and charge distribution. KIV-2 affinity for predicted small molecule binders was found to be negligible in surface plasmon resonance experiments; and with the LBS being nonfunctional, we propose to rename it "pseudo-LBS". Further investigation of the protein by computational small-molecule docking allowed us to identify a possible heparin-binding site away from the LBS, which was confirmed by specific reverse charge mutations abolishing heparin binding. This study opens new possibilities to define the pathogenesis of Lp(a)-related diseases and to facilitate the design of specific therapeutic drugs.
- Department of Molecular Medicine, Unit of Immunology and General Pathology, University of Pavia, Pavia, Italy.
Organizational Affiliation: