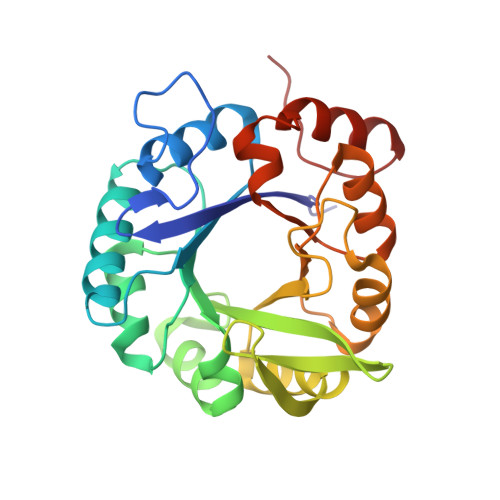
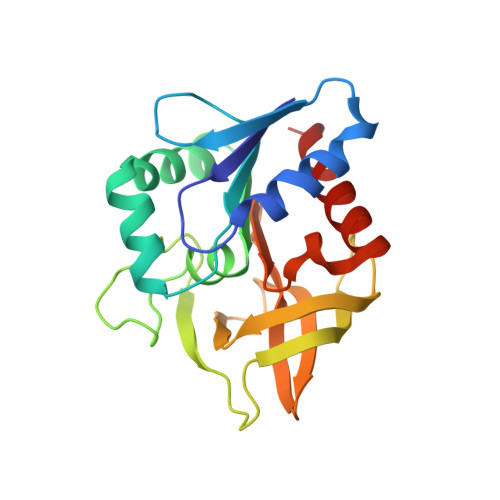
Light Regulation of Enzyme Allostery through Photo-responsive Unnatural Amino Acids.
Kneuttinger, A.C., Straub, K., Bittner, P., Simeth, N.A., Bruckmann, A., Busch, F., Rajendran, C., Hupfeld, E., Wysocki, V.H., Horinek, D., Konig, B., Merkl, R., Sterner, R.(2019) Cell Chem Biol 26: 1501-1514.e9
- PubMed: 31495713
- DOI: https://doi.org/10.1016/j.chembiol.2019.08.006
- Primary Citation Related Structures:
6RTZ, 6RU0 - PubMed Abstract:
Imidazole glycerol phosphate synthase (ImGPS) is an allosteric bienzyme complex in which substrate binding to the synthase subunit HisF stimulates the glutaminase subunit HisH. To control this stimulation with light, we have incorporated the photo-responsive unnatural amino acids phenylalanine-4'-azobenzene (AzoF), o-nitropiperonyl-O-tyrosine (NPY), and methyl-o-nitropiperonyllysine (mNPK) at strategic positions of HisF. The light-mediated isomerization of AzoF at position 55 (fS55AzoF E ↔ fS55AzoF Z ) resulted in a reversible 10-fold regulation of HisH activity. The light-mediated decaging of NPY at position 39 (fY39NPY → fY39) and of mNPK at position 99 (fK99mNPK → fK99) led to a 4- to 6-fold increase of HisH activity. Molecular dynamics simulations explained how the unnatural amino acids interfere with the allosteric machinery of ImGPS and revealed additional aspects of HisH stimulation in wild-type ImGPS. Our findings show that unnatural amino acids can be used as a powerful tool for the spatiotemporal control of a central metabolic enzyme complex by light.
- Institute of Biophysics and Physical Biochemistry, University of Regensburg, Universitätsstrasse 31, 93053 Regensburg, Germany.
Organizational Affiliation: