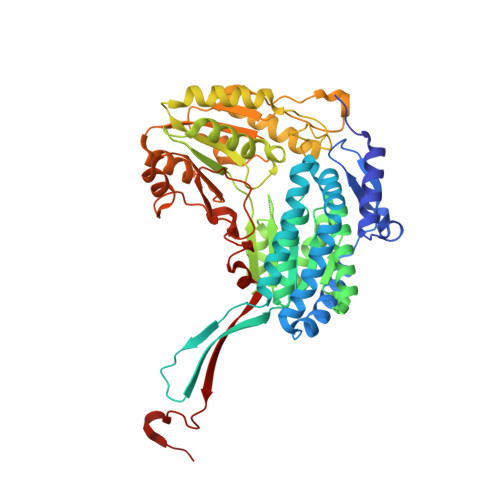
Structure and mechanism of piperideine-6-carboxylate dehydrogenase from Streptomyces clavuligerus.
Hasse, D., Hulsemann, J., Carlsson, G.H., Valegard, K., Andersson, I.(2019) Acta Crystallogr D Struct Biol 75: 1107-1118
- PubMed: 31793904
- DOI: https://doi.org/10.1107/S2059798319014852
- Primary Citation Related Structures:
6RTR, 6RTS, 6RTT, 6RTU - PubMed Abstract:
The core of β-lactam antibiotics originates from amino acids of primary metabolism in certain microorganisms. β-Lactam-producing bacteria, including Streptomyces clavuligerus, synthesize the precursor of the amino acid α-aminoadipic acid by the catabolism of lysine in two steps. The second reaction, the oxidation of piperideine-6-carboxylate (or its open-chain form α-aminoadipate semialdehyde) to α-aminoadipic acid, is catalysed by the NAD + -dependent enzyme piperideine-6-carboxylate dehydrogenase (P6CDH). This structural study, focused on ligand binding and catalysis, presents structures of P6CDH from S. clavuligerus in its apo form and in complexes with the cofactor NAD + , the product α-aminoadipic acid and a substrate analogue, picolinic acid. P6CDH adopts the common aldehyde dehydrogenase fold, consisting of NAD-binding, catalytic and oligomerization domains. The product binds in the oxyanion hole, close to the catalytic residue Cys299. Clear density is observed for the entire cofactor, including the nicotinamide riboside, in the binary complex. NAD + binds in an extended conformation with its nicotinamide ring overlapping with the binding site of the carboxylate group of the product, implying that the conformation of the cofactor may change during catalysis. The binding site of the substrate analogue overlaps with that of the product, suggesting that the cyclic form of the substrate, piperideine-6-carboxylate, may be accepted as a substrate by the enzyme. The catalytic mechanism and the roles of individual residues are discussed in light of these results.
- Laboratory of Molecular Biophysics, Department of Cell and Molecular Biology, Uppsala University, Husargatan 3, Box 596, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: