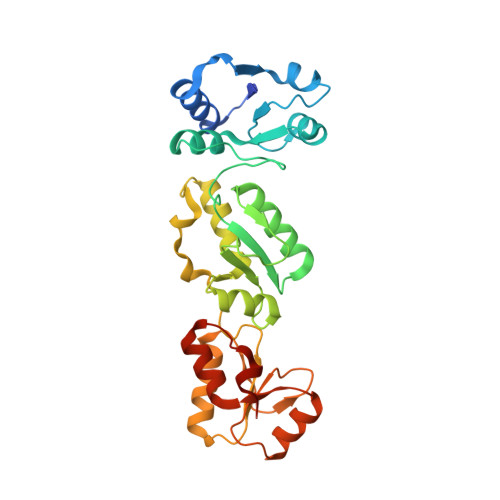
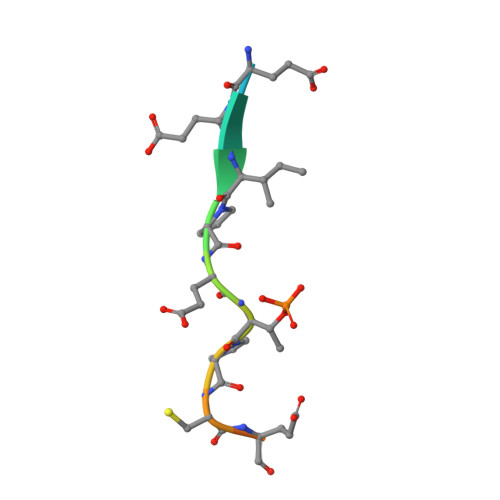
Phosphorylation-mediated interactions with TOPBP1 couple 53BP1 and 9-1-1 to control the G1 DNA damage checkpoint.
Bigot, N., Day, M., Baldock, R.A., Watts, F.Z., Oliver, A.W., Pearl, L.H.(2019) Elife 8
- PubMed: 31135337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.44353
- Primary Citation Related Structures:
6RML, 6RMM - PubMed Abstract:
Coordination of the cellular response to DNA damage is organised by multi-domain 'scaffold' proteins, including 53BP1 and TOPBP1, which recognise post-translational modifications such as phosphorylation, methylation and ubiquitylation on other proteins, and are themselves carriers of such regulatory signals. Here we show that the DNA damage checkpoint regulating S-phase entry is controlled by a phosphorylation-dependent interaction of 53BP1 and TOPBP1. BRCT domains of TOPBP1 selectively bind conserved phosphorylation sites in the N-terminus of 53BP1. Mutation of these sites does not affect formation of 53BP1 or ATM foci following DNA damage, but abolishes recruitment of TOPBP1, ATR and CHK1 to 53BP1 damage foci, abrogating cell cycle arrest and permitting progression into S-phase. TOPBP1 interaction with 53BP1 is structurally complimentary to its interaction with RAD9-RAD1-HUS1, allowing these damage recognition factors to bind simultaneously to the same TOPBP1 molecule and cooperate in ATR activation in the G1 DNA damage checkpoint.
- Cancer Research UK DNA Repair Enzymes Group, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Organizational Affiliation: