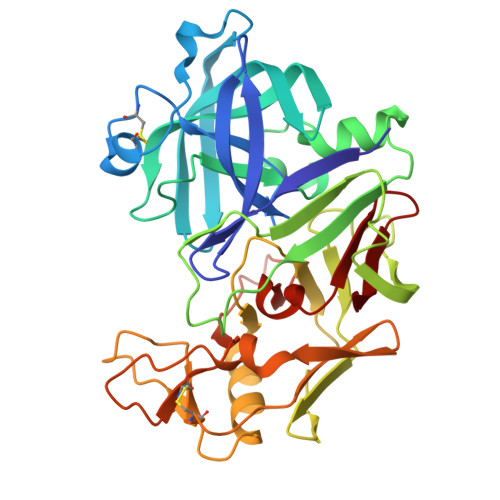
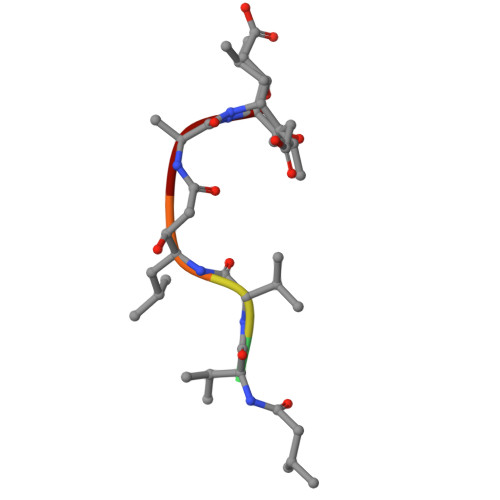
Re-emerging Aspartic Protease Targets: Examining Cryptococcus neoformans Major Aspartyl Peptidase 1 as a Target for Antifungal Drug Discovery.
Krystufek, R., Sacha, P., Starkova, J., Brynda, J., Hradilek, M., Tloust'ova, E., Grzymska, J., Rut, W., Boucher, M.J., Drag, M., Majer, P., Hajek, M., Rezacova, P., Madhani, H.D., Craik, C.S., Konvalinka, J.(2021) J Med Chem 64: 6706-6719
- PubMed: 34006103
- DOI: https://doi.org/10.1021/acs.jmedchem.0c02177
- Primary Citation of Related Structures:
6R5H, 6R6A - PubMed Abstract:
Cryptococcosis is an invasive infection that accounts for 15% of AIDS-related fatalities. Still, treating cryptococcosis remains a significant challenge due to the poor availability of effective antifungal therapies and emergence of drug resistance. Interestingly, protease inhibitor components of antiretroviral therapy regimens have shown some clinical benefits in these opportunistic infections. We investigated Major aspartyl peptidase 1 (May1), a secreted Cryptococcus neoformans protease, as a possible target for the development of drugs that act against both fungal and retroviral aspartyl proteases. Here, we describe the biochemical characterization of May1, present its high-resolution X-ray structure, and provide its substrate specificity analysis. Through combinatorial screening of 11,520 compounds, we identified a potent inhibitor of May1 and HIV protease. This dual-specificity inhibitor exhibits antifungal activity in yeast culture, low cytotoxicity, and low off-target activity against host proteases and could thus serve as a lead compound for further development of May1 and HIV protease inhibitors.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Flemingovo n. 2, Prague 6 16610, Czech Republic.
Organizational Affiliation: