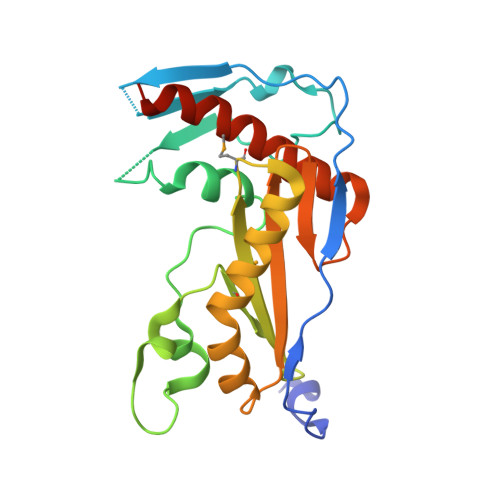
Structure and function of the Toscana virus cap-snatching endonuclease.
Jones, R., Lessoued, S., Meier, K., Devignot, S., Barata-Garcia, S., Mate, M., Bragagnolo, G., Weber, F., Rosenthal, M., Reguera, J.(2019) Nucleic Acids Res 47: 10914-10930
- PubMed: 31584100 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz838
- Primary Citation Related Structures:
6QVV, 6QW0, 6QW5 - PubMed Abstract:
Toscana virus (TOSV) is an arthropod-borne human pathogen responsible for seasonal outbreaks of fever and meningoencephalitis in the Mediterranean basin. TOSV is a segmented negative-strand RNA virus (sNSV) that belongs to the genus phlebovirus (family Phenuiviridae, order Bunyavirales), encompassing other important human pathogens such as Rift Valley fever virus (RVFV). Here, we carried out a structural and functional characterization of the TOSV cap-snatching endonuclease, an N terminal domain of the viral polymerase (L protein) that provides capped 3'OH primers for transcription. We report TOSV endonuclease crystal structures in the apo form, in complex with a di-ketoacid inhibitor (DPBA) and in an intermediate state of inhibitor release, showing details on substrate binding and active site dynamics. The structure reveals substantial folding rearrangements absent in previously reported cap-snatching endonucleases. These include the relocation of the N terminus and the appearance of new structural motifs important for transcription and replication. The enzyme shows high activity rates comparable to other His+ cap-snatching endonucleases. Moreover, the activity is dependent on conserved residues involved in metal ion and substrate binding. Altogether, these results bring new light on the structure and function of cap-snatching endonucleases and pave the way for the development of specific and broad-spectrum antivirals.
- Aix-Marseille Université, CNRS, AFMB UMR 7257, 13288 Marseille, France.
Organizational Affiliation: