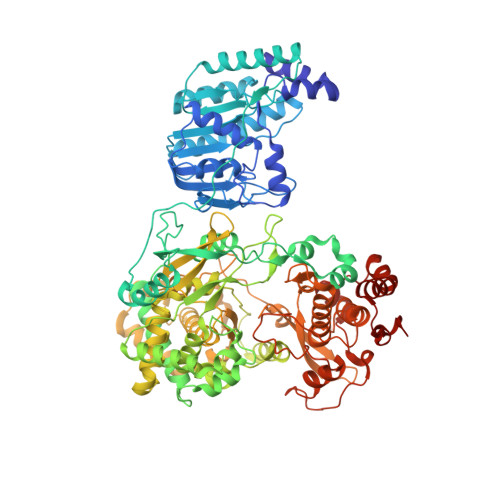
Structure of the yellow fever NS5 protein reveals conserved drug targets shared among flaviviruses.
Dubankova, A., Boura, E.(2019) Antiviral Res 169: 104536-104536
- PubMed: 31202975
- DOI: https://doi.org/10.1016/j.antiviral.2019.104536
- Primary Citation Related Structures:
6QSN - PubMed Abstract:
Yellow fever virus (YFV) is responsible for devastating outbreaks of Yellow fever (YF) in humans and is associated with high mortality rates. Recent large epidemics and epizootics and exponential increases in the numbers of YF cases in humans and non-human primates highlight the increasing threat YFV poses, despite the availability of an effective YFV vaccine. YFV is the first human virus discovered, but the structures of several of the viral proteins remain poorly understood. Here we report the structure of the full-length NS5 protein, a key enzyme for the replication of flaviviruses that contains both a methyltransferase domain and an RNA dependent RNA polymerase domain, at 3.1 Å resolution. The viral polymerase adopts right-hand fold, demonstrating the similarities of the Yellow fever, Dengue and Zika polymerases. Together this data suggests NS5 as a prime and ideal target for the design of pan-flavivirus inhibitors.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Flemingovo Nam. 2, 166 10, Prague 6, Czech Republic.
Organizational Affiliation: